Personalizing Breast Cancer Treatments by Targeting Specific Proteins
One of the major aims of biomedical research is to improve patient therapy, and as more powerful research and clinical tools have become available, therapeutics that are tailored to individual patients has become a more realistic goal. Scientists have now developed a method for a more personalized approach to breast cancer treatment, by both targeting the proteins that help drive tumor growth and evaluating the efficacy of drug treatments that act upon those drivers. The work, which was reported in Nature Communications, is summarized in the following video.
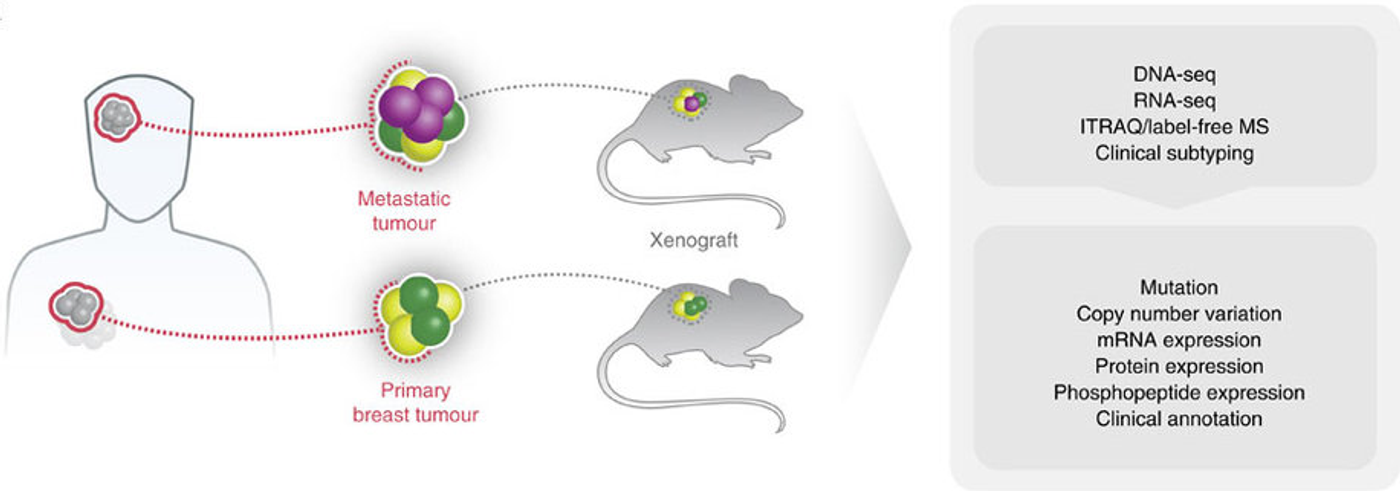
"Here we study the problem of how to design more effective cancer treatments with a two-pronged approach," commented one co-author of the report, Dr. Matthew Ellis, Professor and Director of the Lester and Sue Smith Breast Cancer Center at Baylor. "We combine patient-derived xenografts and proteogenomic integration." Patient-derived [tumor] xenografts are created by implanting a sample taken from a human cancer tumor into a lab mouse, specifically one that is immunodeficient to minimize transplant rejection.
"We grow human tumors direct from patients in laboratory mice because we want to mimic the living natural environment tumor cells grow in - a beating heart, blood vessels and other biological structures - that is a natural as we can possibly make it," explained Ellis. "We think this approach offers a closer representation of the tumor's growth environment to study cancer drugs than cells growing on a laboratory plastic dish."
It’s known that cancer cells express different genes than normal cells do, which leads to differential protein expression as well. Previous research has demonstrated that this method creates a good model; patient-derived xenografts of breast cancer tumors demonstrate a gene and protein expression profile that is like that seen in the original breast tumors. In addition, these xenografts seem to react to drug treatments on ways that are similar to what is observed in the tumors of human patients.
"Proteins carry out most of the biological functions in the cell," said Dr. Li Ding, an Associate Professor of Medicine at Washington University School of Medicine. "Knowing the DNA sequence of genes does not automatically tell us everything about the proteins doing work in the cells."
For this work, the investigators utilized mass spectrometry, which is a common technique that can quantify and characterize proteins to identify the proteins present in 24 different patient-derived xenografts of breast tumors of various subtypes. The research team found and measured the levels of 10,000 proteins in each of the tumors.
The next step was to test therapeutics; in some patient-derived xenografts of breast tumors specific dysfunctional proteins were targeted by specific inhibitors. The researchers were successful; they observed a reduction of tumor growth in the disease model.
"A substantial number of these proteins identified in the xenograft model are potential targets for drugs," Ellis said. "In addition, similar protein signatures were observed in breast cancer samples from patients, which suggests our approach has potential clinical relevance."
He expanded on how this idea can be applied to personalized medicine. “Eventually, with this new approach scientists will be able to answer questions such as why a certain drug that induced one tumor to shrink in one patient did not affect the growth of a similar tumor in another patient,” Ellis said. “The idea behind this approach is to find out what are the driving pathways of each person’s cancer. Once we know these pathways, the next step is to use drugs to interrupt these pathways and lead to outcomes that are better than giving patients non-specific therapy. The mouse work is promising enough to adapt these technologies for real time analysis of patient samples so that clinical trials can be designed to test this new diagnostic and drug selection approach. The National Cancer Institute has agreed to invest in this next phase of the project.”
Sources: UPI, Baylor College of Medicine, Nature Communications