Precise Visualization of CRISPR Tool Aims to Improve Accuracy
The CRISPR gene-editing tool has been widely adopted since its relatively recent invention, and researchers are still working to learn more about all the details surrounding both its mode of action and potential off-target effects that could be deleterious to the system. New work is sure to help refine the technique; it has provided insight into the mechanism through snapshots that are nearly atomic level. The research by scientists at Harvard Medical School and Cornell University has been published in Cell.
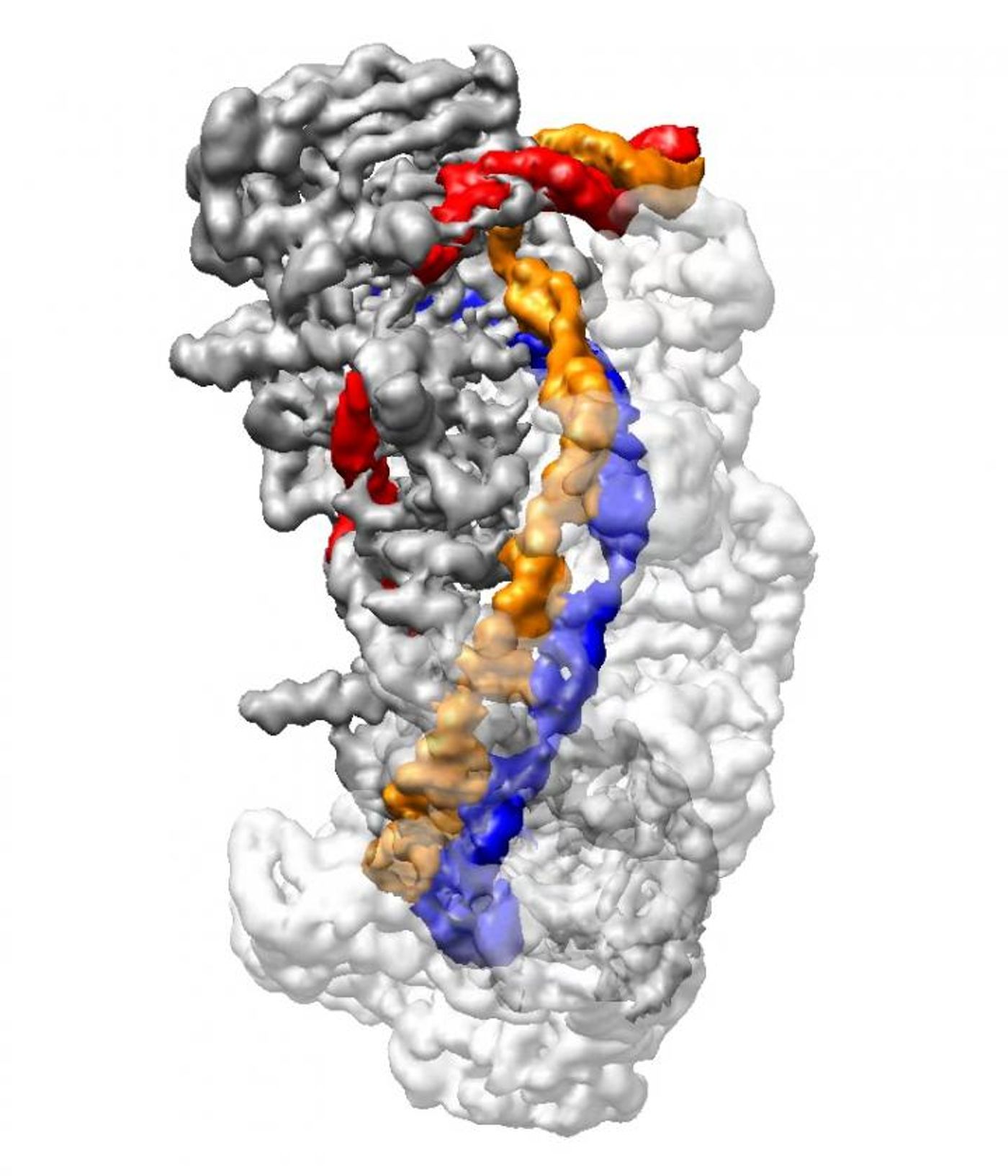
Cryo-electron microscopy allowed the scientists to elucidate the process behind CRISPR-Cas3, a variant on the CRISPR-Cas9 DNA editing system. This technique enables a genomic edit, while ensuring multiple levels of error detection that aims to prevent damage to the genome. This new data will help improve CRISPR accuracy by providing structural imagery of CRISPR machinery interacting with its target DNA and getting it ready for a cut by the Cas3 enzyme.
Scientists use the CRISPR-Cas system, especially Cas9, pairing with RNA sequences generated in the lab and enabling the precise editing of genomic material with unprecedented simplicity.
This work is an important step forward in use of the tool in human applications. "To solve problems of specificity, we need to understand every step of CRISPR complex formation," said co-senior author Maofu Liao, an Assistant Professor of Cell Biology at Harvard Medical School. "Our study now shows the precise mechanism for how invading DNA is captured by CRISPR, from initial recognition of target DNA and through a process of conformational changes that make DNA accessible for final cleavage by Cas3."
CRISPR-Cas is naturally used by bacteria for defense. Snippets of viral DNA get taken up by bacteria and integrated into its genome . Short RNA sequences can subsequently be made; they are called crRNA (CRISPR RNA). These crRNAs are used to identify enemy invaders. The crRNA is bound to various CRISPR enzymes, which then act as monitors for those invaders. When these riboprotein machines find something that matches its crRNA, they destroy the DNA.
Liao collaborated with Ailong Ke of Cornell University for an investigation of type 1 CRISPR, which uses a riboprotein complex called CRISPR Cascade for the capture of target DNA and the Cas3 enzyme to make DNA cuts. With CRISPR-Cas3, crRNA joins with CRISPR Cascade, which looks for PAM, a very short DNA sequence that indicates that foreign viral DNA is present.
Liao, Ke and colleagues determined that when Cascade finds PAM, it bends DNA sharply at an angle, and forces a bit of the DNA to unwind. The crRNA can then bind a short strand of target DNA, creating a "seed bubble."
This seed bubble functions to ensure that target DNA matches the crRNA. If there is a correct match, the bubble enlarges, and the rest of the crRNA is then bound to its intended target DNA. This forms an "R-loop" structure.
After completing the formation of the R-loop, the CRISPR Cascade complex changes its shape, and locks the DNA in place. In the second, non-targeted strand of DNA another bulge is made.
The Cas3 enzyme binds and cuts only after the formation of a full R-loop state is generated, showing that there are elaborate checks on the system. In nature, this ensures that bacteria will not accidentally destroy its own DNA.
"To apply CRISPR in human medicine, we must be sure the system is accurate and that it does not target the wrong genes," said Ke, the co-senior author of the study. "Our argument is that the CRISPR-Cas3 subtype has evolved to be a precise system that carries the potential to be a more accurate system to use for gene editing. If there is mis-targeting, we know how to manipulate the system because we know the steps involved and where we might need to intervene."
This is the first look at the full process, from the formation of a seed bubble to the R-loop at extremely high resolution. Contrasting with the scalpel-like edit of Cas9, Cas3 acts more like a shredder, which makes it better suited for use against antibiotic-resistant bacteria.
Additionally, CRISPR-Cas variants all use some kind of R-loop formation for the detection and preparation of target DNA for cuts. This new structural understanding can help scientists modify other kinds of CRISPR-Cas systems, thereby reducing the potential for undesired off-target effects.
"Scientists hypothesized that these states existed but they were lacking the visual proof of their existence," said co-first author Min Luo, postdoctoral fellow in the Liao lab at HMS. "The main obstacles came from stable biochemical reconstitution of these states and high-resolution structural visualization. Now, seeing really is believing."
"We've found that these steps must occur in a precise order," Luo said. "Evolutionarily, this mechanism is very stringent and has triple redundancy, to ensure that this complex degrades only invading DNA."
A review of CRISPR:
Sources: AAAS/Eurekalert! via Harvard Medical School, Cell