Changing our View of DNA Organization
The DNA from a single human cell can stretch about five feet; the DNA from all of the cells in our bodies could stretch to Pluto! All of that genetic material has to be carefully organized so it can fit into the cell nucleus, a tiny organelle that usually spans about a thousandth of a millimeter. Our cells also have to be able to access specific regions of the genome when needed, so the organization has to be carefully controlled. Now, scientists at the Salk Institute have revealed an amazing, three-dimensional look at chromatin, which is DNA in a more natural state, with all of its associated proteins attached.
Reporting in Science, researchers have paired a new dye with powerful microscopy techniques to visualize chromatin in cells at rest, and as cell division is happening. These findings could rewrite textbooks. Up until now, chromatin has been studied after harsh treatment in the lab. This work allowed investigators to get a look at it as it exists in nature. Learn more from the video.
"One of the most intractable challenges in biology is to discover the higher-order structure of DNA in the nucleus and how is this linked to its functions in the genome," said senior author Clodagh O'Shea, a Howard Hughes Medical Institute Faculty Scholar and Salk Associate Professor. "It is of eminent importance, for this is the biologically relevant structure of DNA that determines both gene function and activity."
Understanding the exact details of DNA structure has eluded scientists. It has been thought that DNA is spooled around proteins, forming nucleosomes that are sometimes likened to beads on a string. Those 11-nanometer beads are thought to fold up into fibers that get increasingly bigger, eventually forming chromosomes. The intermediate fiber structure has not been seen directly in a living cell, however, only after laboratory processing.
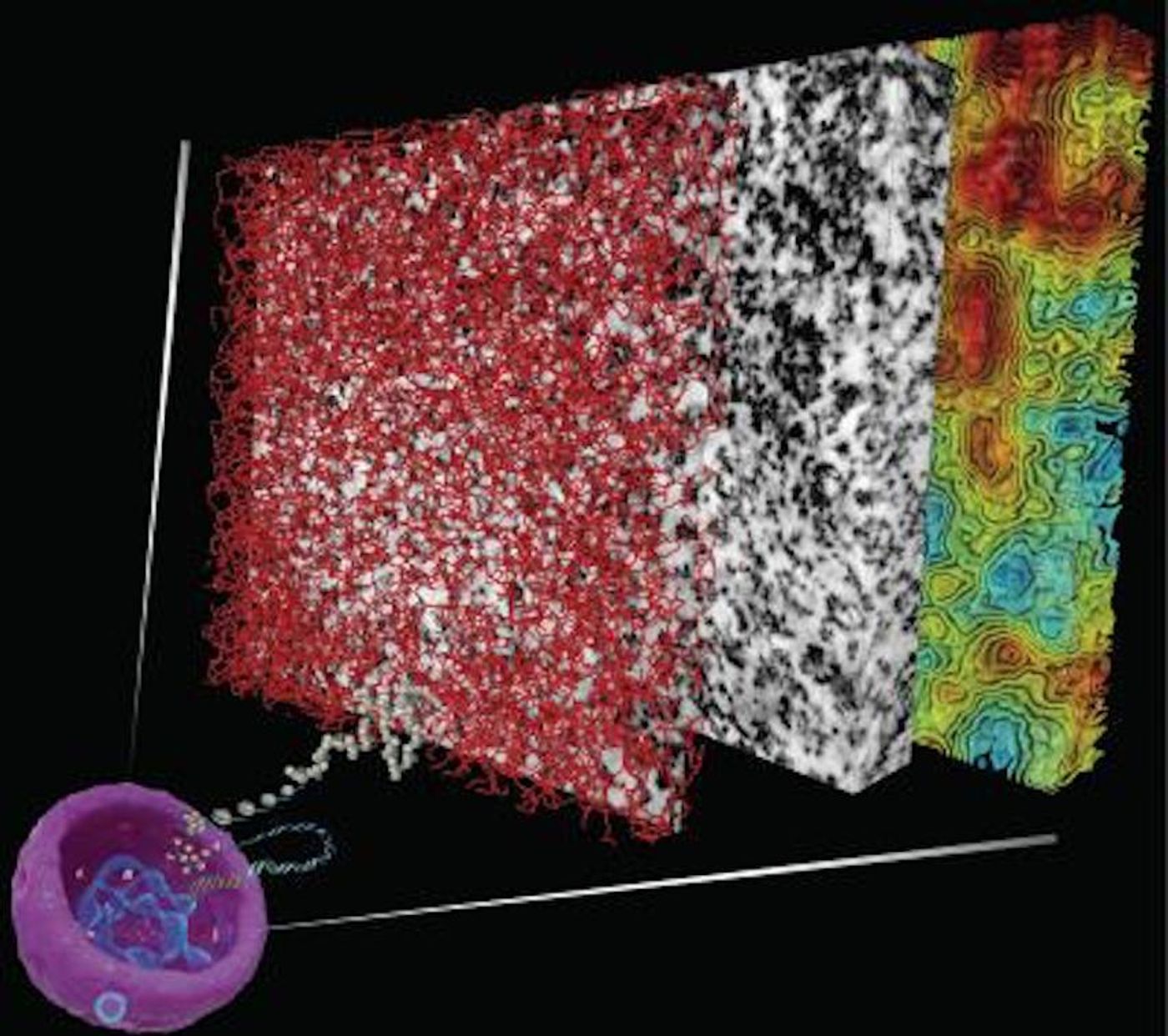
For this work, the researchers wanted to see the chromatin in an untreated nucleus, and eventually found a dye that could coat DNA with a metal, allowing visualization of structure and organization at an unprecedented level. Another author of the work, microscopy expert Mark Ellisman, a Professor at the University of California, San Diego, utilized a powerful form of electron microscopy and combined it with the chromatin dye to create electron-microscope tomography, ChromEMT.
The scientists used ChromEMT to analyze chromatin in human cells in various states - at rest, and when compacted during cell division. They were surprised by their data; they never observed the secondary structures described in textbooks.
"The textbook model is a cartoon illustration for a reason," noted first author Horng Ou, a Salk Institute research associate. "Chromatin that has been extracted from the nucleus and subjected to processing in vitro--in test tubes--may not look like chromatin in an intact cell, so it is tremendously important to be able to see it in vivo."
Instead, the researchers saw that in during both division and rest, the beads on a string never folded into the fibers. It formed a chain that could bend and flex, with varying levels of density. The researchers carefully took measure, finding that it had a length of between five and 24 nanometers. These findings suggest that packing density, and not secondary structure, could be relevant to which areas of the genome become active or inactive.
"We show that chromatin does not need to form discrete higher-order structures to fit in the nucleus," explained O'Shea. "It's the packing density that could change and limit the accessibility of chromatin, providing a local and global structural basis through which different combinations of DNA sequences, nucleosome variations, and modifications could be integrated in the nucleus to exquisitely fine-tune the functional activity and accessibility of our genomes."
This work could have implications for disease treatment if scientists can gain a better understanding of how to control the genome by manipulating chromatin or access to it. Next, scientists want to see if the chromatin structure they observed is common among cell types and organisms.
Sources: AAAS/Eurkealert! Via Salk Institute, Science