Chipping Away at the Mystery of Viruses
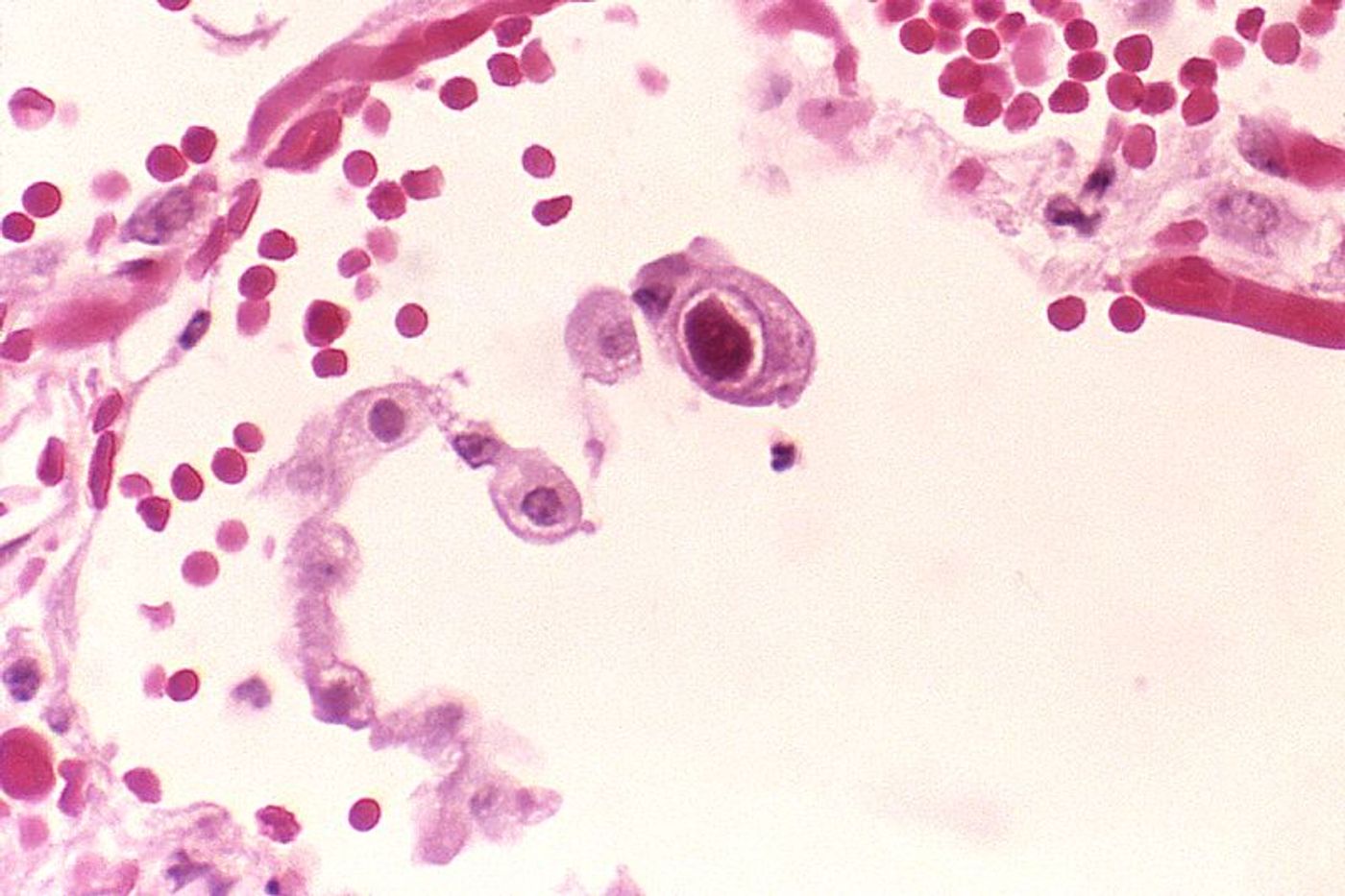
If an adult gets infected with cytomegalovirus, it usually doesn’t cause any harm, unless that adult is pregnant. In those cases, it's possible for the virus to infect a developing fetus, which can result in birth defects. Now researchers have completed an analysis of the more than 500 viral proteins and peptides that are made after the virus invades a cell. A new bioinformatics tool has revealed this protein profile, which includes 200 proteins that were completely unknown before this.
Researchers at the Department of Virology of Julius-Maximilians-Universität Würzburg (JMU) in Bavaria, Germany believe that their bioinformatics technique, called PRICE, will have applications in medicine; an understanding of viral proteins may enable scientists to create therapeutics and vaccines for viruses.
Reporting in Nature Methods, the teams of Professor Lars Dölken and Professor Florian Erhard have created a way to accurately capture the behavior of a cell’s ribosomes, organelles that perform the critical work of assembling proteins. A virus simply hijacks the machinery a cell already contains, so those ribosomes are used to manufacture viral proteins after infection. Molecules called mRNAs take the instructions for assembly from genes to the organelle, where they are translated into proteins.
There are many questions about the manufacture of viral proteins after a cell is infected. An important technique, ribosomal profiling or Ribo-seq, can help answer those questions. Ribosomal activity, or translation processes, are assessed by Ribo-seq as patterns. PRICE is a better way to identify translation events.
"Previously, a number of error sources often prevented the reliable detection of translation events when analyzing Ribo-seq data," said Erhard. In mRNA, sequences called shorter open reading frames (sORFs) sometimes come before open reading frames (ORFs), which are well known.
"Our method is capable of resolving even complex cases, for example overlapping ORFs or unusual start codons, with high accuracy. This has allowed us to determine all translated areas genome-wide with high accuracy for the first time," Professor Erhard explained.
The tool allowed the JMU team to find many new cellular and viral proteins and peptides. They also found that many sORF peptides are used on the surface of cells by MHC-1 molecules.
"sORFs thus encode for a new class of antigens that can be recognized by our immune system," said Dölken. "We therefore assume that sORFs are involved in immunological control mechanisms especially during virus infections and stress responses." This work opens new avenues for the investigation of viral infections.
"PRICE now enables us to analyze all existing and future dataset in much greater depth and with substantially improved accuracy," added Dölken. He suggested that it may be worth the effort to take a second look at any data that has already been published. To encourage that effort, the analysis tools have been made freely accessible online by the scientists.
Sources: AAAS/Eurekalert! Via University of Würzburg, Nature Methods