A Brain Map Made by Crowdsourcing
Princeton researchers turned to gamers when they needed to make sense of a huge amount of data that was created to understand more about cells in the brain. About a quarter of a million game players worked as part of Eyewire to create interactive maps of thousands of neurons, now freely available to neuroscientists and the public, by the Brain Research through Advancing Innovative Neurotechnologies (BRAIN) Initiative.
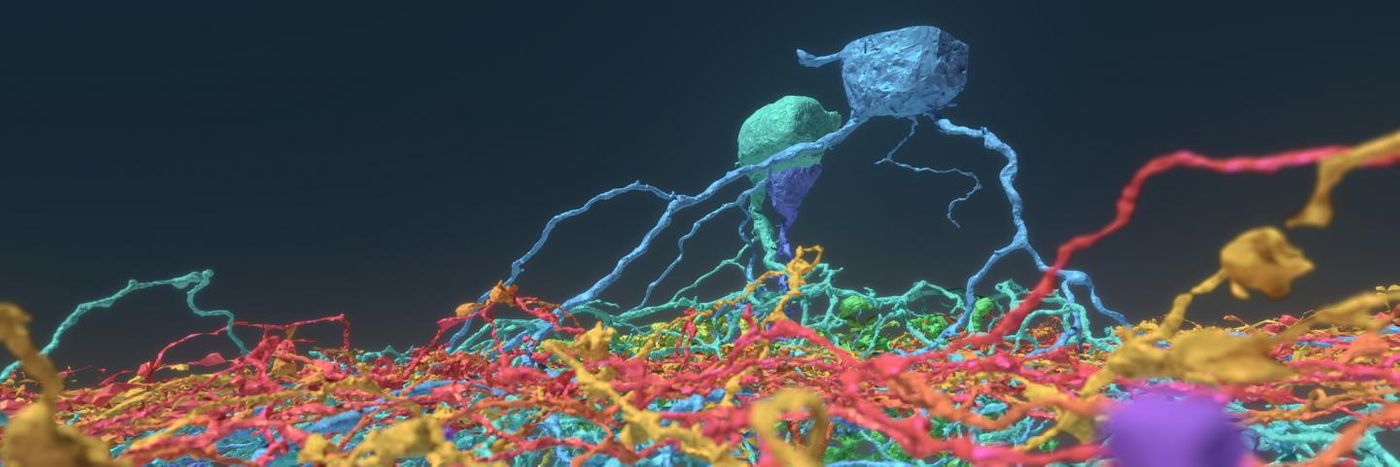
"Working with Eyewirers around the world, we've made a digital museum that shows off the intricate beauty of the retina's neural circuits," said Sebastian Seung, the Evnin Professor in Neuroscience and a professor of computer science and the Princeton Neuroscience Institute (PNI). The Eyewire museum was unveiled by Seung, and the work has been reported in Cell.
"This interactive viewer is a huge asset for these larger collaborations, especially among people who are not physically in the same lab," noted Amy Robinson Sterling, a crowdsourcing specialist with PNI and the executive director of Eyewire, the gaming platform used online by citizen scientists who helped make the viewer.
"This museum is something like a brain atlas," said one of the co-first authors, Alexander Bae, a graduate student in electrical engineering. "Previous brain atlases didn't have a function where you could visualize by individual cell, or a subset of cells, and interact with them. Another novelty: Not only do we have the morphology of each cell, but we also have the functional data, too."
With data from a mouse retina study done in 2009, neural maps were created by the Eyewirers. Machine learning was used in conjunction with manual work by the gamers, who were better equipped than computers to trace the path of every twisting neuron. Patterns of neurons were recorded, and the data was checked by other advanced players, staff members at Eyewire, and self-improving pattern recognition software. Eyewire uses 3D cubes which are a subset of one cell; five to 25 gamers reviewed each cube. There were over ten million of those cubes.
"Back in the early years it took weeks to finish a single cell," said Sterling. "Now players complete multiple neurons per day." The Eyewire user experience stays focused on the larger mission - "For science!" is a common refrain - but it also replicates a typical gaming environment, with achievement badges, a chat feature to connect with other players and technical support, and the ability to unlock privileges with increasing skill. "Our top players are online all the time, easily 30 hours a week," Sterling said.
The gameplay has been made customizable and promoted to make it more fun. "The community has really been the driving force behind why Eyewire has been successful," Sterling said. "You come in, and you're not alone. Right now, there are 43 people online. Some of them will be admins from Boston or Princeton, but most are just playing -- now it's 46."
The researchers focused on retinal ganglion cells, which will help reveal how the retina functions. The cells grow from the same developmental tissue that the brain originates from.
"The retina doesn't just sense light," Seung said. "Neural circuits in the retina perform the first steps of visual perception."
The video explains how to play Eyewire.
"Why the ganglion cells of the eye?" asked Shang Mu, an associate research scholar in PNI and fellow first author. "Because they're the connection between the retina and the brain. They're the only cell class that go back into the brain."
The study has found six new types of cells; a literature search did not reveal any previous reports of their existence. "The number of different cell types was a surprise," said Mu. "Just a few years ago, people thought there were only 15 to 20 ganglion cell types, but we found more than 35 - we estimate between 35 and 50 types."
Sources: AAAS/Eurekalert! Via Princeton, Cell