Analyzing Genomic Data in the Field
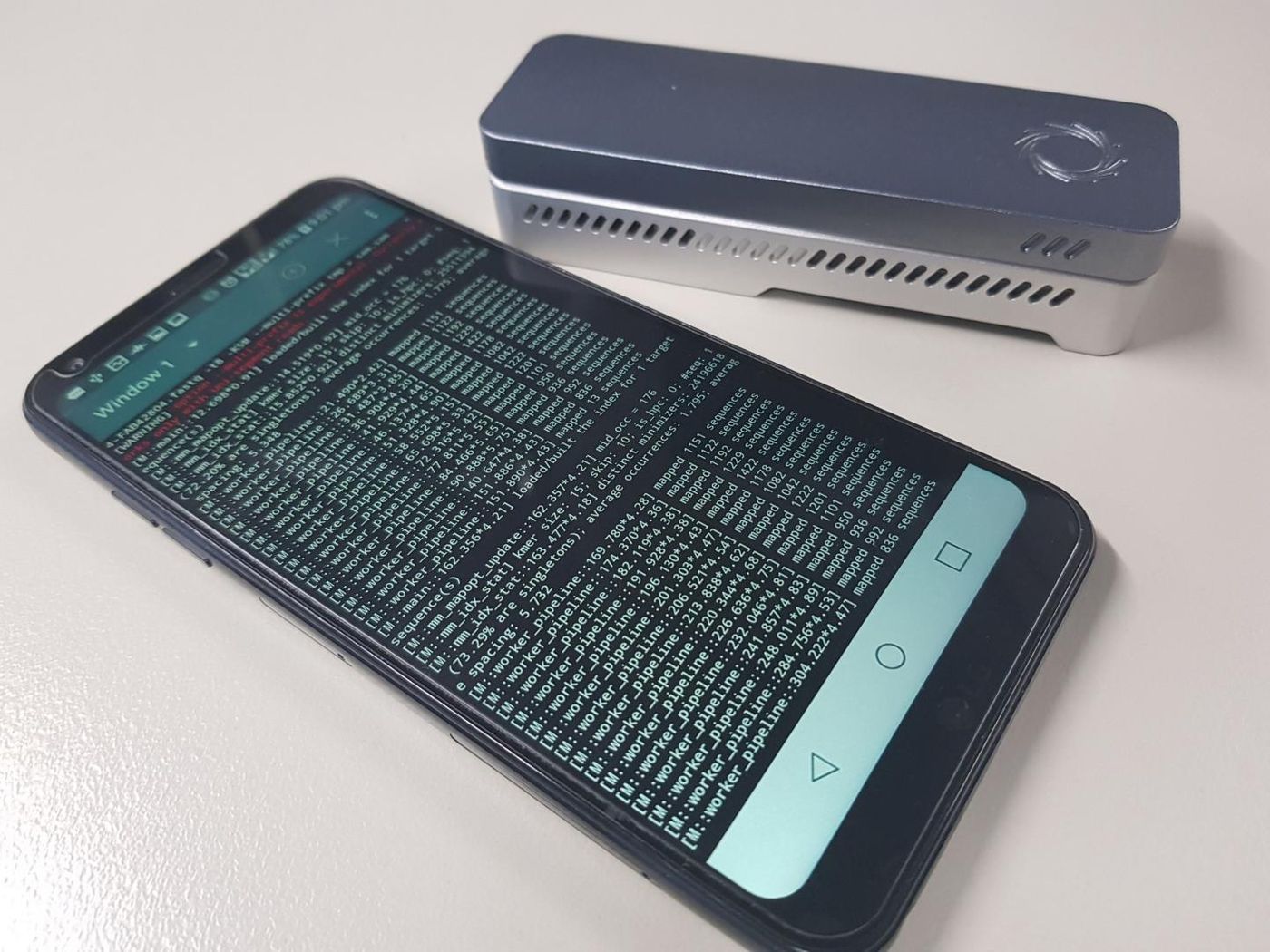
Genetic technologies have advanced rapidly in recent years, and healthcare is moving closer to a time when doctors will consult genetic sequencing data when advising patients. Devices that can sequence whole genomes are getting smaller and require less computational power, like the MinION Sequencer by Oxford Nanopore Technologies. But they still need to be hooked up to a larger computer or internet connection in order to analyze it, by comparing it to a reference genome.
Now, scientists at the Garvan Institute of Medical Research and UNSW Sydney have developed a new method for taking that analysis offline. They created an algorithm that needs only 2GB of memory to align genomic sequences, instead of the 16GB that used to be necessary. Now genomic analysis can be done in the field, with the computational power of a smartphone. The work has been published in Scientific Reports.
"We're focused on making genomic technologies more accessible to improve human health. They're becoming smaller, but still need to function in remote areas, so we created a method that can analyze genomic data, in real time, on just a mobile device," said Dr. Martin Smith. He is the Team Leader of Genomic Technologies at the Garvan Institute's Kinghorn Centre for Clinical Genomics.
The team utilized Minimap2, a program that aligns lengths of DNA sequencing reads to known reference genomes. The references can be indexed, so the reads get quickly mapped to their places in a reference genome. Handling the references and aligning reads to them takes power.
"The challenge, so far, has been that the reference index requires too much computer memory. We took the approach of splitting the reference library up into smaller segments, against which we mapped the DNA reads. Once we finished mapping to the smaller segments, we pool results together and tease out the noise, much like creating a panorama by stitching together smaller photos," Smith explained.
"Other algorithms, which take a similar approach of splitting up the reference data, produce a lot of spurious and duplicate mappings - just like overlapping photos in the panorama. What we did in this study was fine-tune parameters and select the best mappings across several small indexes. This approach gave us similar accuracy as current standard genomic analyses, which previously required the memory available in high-performance computers," he added.
The researchers validated their algorithm by checking its results against standard tools. The results were replicated 99.98 percent of the time, and with smaller indexes, another one percent could be mapped by the algorithm.
"The potential of lightweight, portable genomic analysis is vast - we hope that this technology will one day be applied in the context of point-of-care microbial infections in remote regions, or in doctors' hands at the hospital bedside," noted Smith.
Learn more about the MinION Sequencer from the video. The device has already been used during Ebola outbreaks and to fight tuberculosis in Madagascar.
Sources: AAAS/Eurekalert! Via Garvan Institute of Medical Research, Scientific Reports