Sparing the Lives of Research Mice with a Gene Activity Database
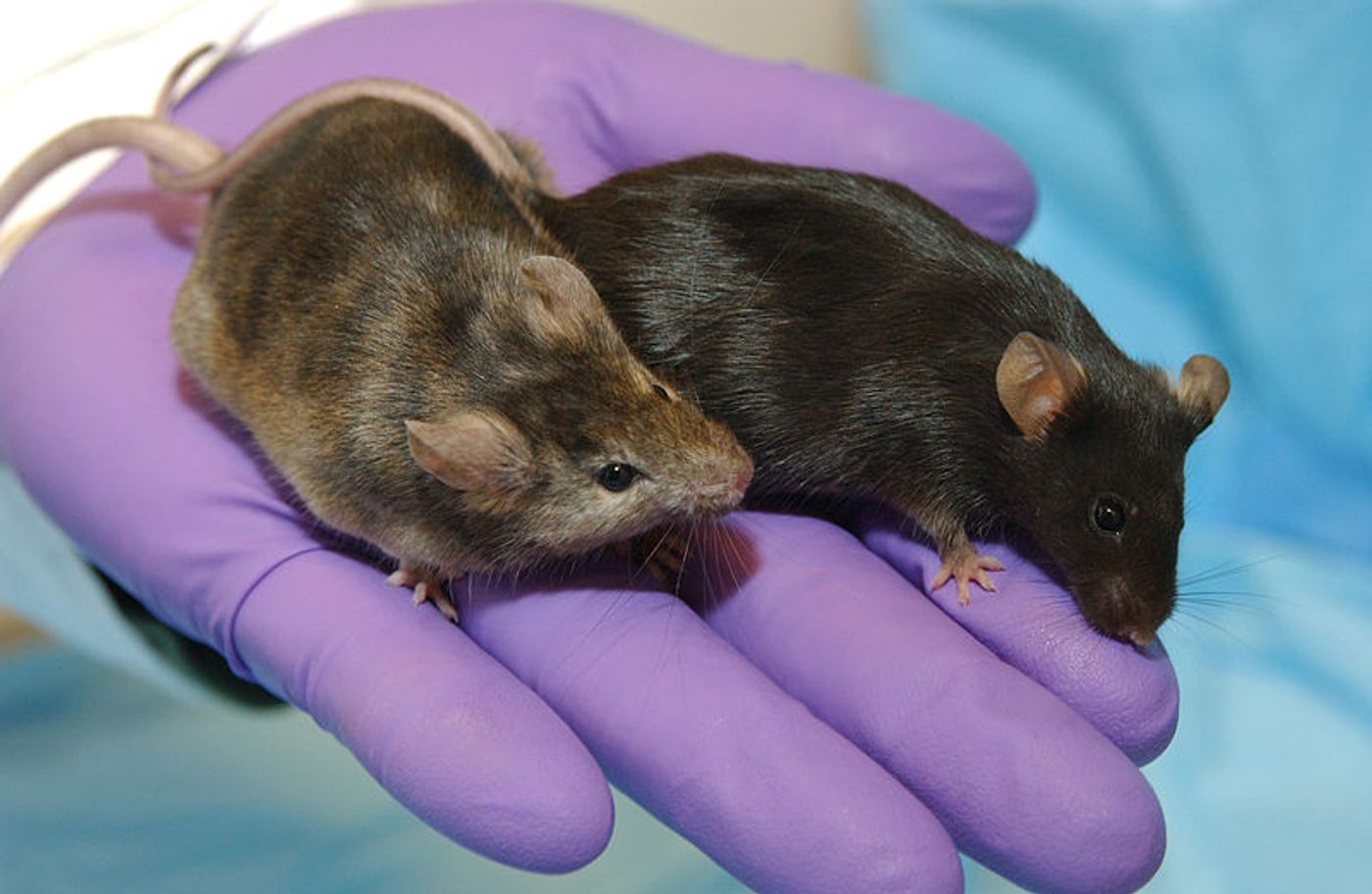
To assess gene activity in a mouse model, scientists have to sacrifice mice to obtain RNA; they can then either focus on the genes they are interested in, or obtain a snapshot of all the active genes. If researchers are modeling an infectious pathogen, for example, they have to infect the mice, then kill them to harvest the necessary materials. Now investigators at the Crick Institute are aiming to reduce the numbers of mice that are needed for such experiments. They've generated a database that catalogues how more than 45,000 mouse genes respond to ten different diseases. Reporting in Nature Communications, the information is accessible with an online app. Learn why researchers use mice from the video.
In this work, blood, as well as lung tissue samples for diseases of the respiratory system, were analyzed using RNA sequencing (RNA-seq). Since active genes are transcribed by cells into RNA, this tool illustrates gene activity, and how it can change because of illness or allergy.
"Gene activity can show us how the body responds to infections and allergens," explained Crick group leader Anne O'Garra. "There are thousands of genes involved in any immune response, so Akul Singhania, a Bioinformatics Postdoc in our lab used advanced bioinformatics approaches to cluster the genes into modules. These modules represent clusters of genes that are co-regulated and can often be annotated to determine their function and known physiological roles. For example, of 38 lung modules there is a module associated with allergy, and seen only in the allergy model, containing over 100 genes and another module associated with T cells containing over 200 genes."
With the app, scientists can explore how various pathogens, like Candida albicans, Toxoplasma gondii parasites, the flu and Respiratory Syncytial Virus, and the bacterium Burkholderia pseudomallei impact the activity of mouse genes in blood and the lungs. Data on gene activity in the blood is available for mice infected with murine cytomegalovirus, listeria, the malaria parasite Plasmodium chabaudi, or a chronic Burkholderia infection.
Information about gene expression in the blood can help show how the immune system is responding, and whether localized responses in the lung reflect it, noted O'Garra. "This will help us to understand what we can learn from genetic signatures in the blood, since for most diseases doctors can't realistically get lung samples from patients."
The different pathogens triggered varied immune responses in the lungs. The scientists were surprised to find that the genes of type I interferons, which respond to viruses, were very active in the blood and lungs of Toxoplasma-infected mice and to a lesser extent, Burholderia bacteria. This indicates that earlier work suggesting that genes associated with type 1 interferons are indicative of viruses is not necessarily true.
"Ten years since the project began, we now have an open access resource of gene expression that anyone in the world can use to look up their favorite genes and also see if they are regulated by type I or type II interferon signalling," said O'Garra. "Nobody said science was easy, but it's certainly worthwhile."
The Jackson Laboratory has also created a powerful tool for exploring and comparing mouse gene expression and relating it to human disease. Learn more about it from the video.
Sources: AAAS/Eurekalert! via The Crick Institute, Nature Communications