Generating Error-Free, Complete Reference Genomes for Vertebrates
Researchers have completed a wave of studies examining the genomes of different vertebrate species. In 20 papers, 25 genomes have been reported from 16 species. To date, researchers have sequenced the genomes of about 400 species, but this work is a major improvement on the quality of those sequences, and even includes mitochondrial genomes. The effort has revealed new chromosomes in the zebra finch and platypus, and has revealed more about the evolution of several species. The research can teach us more about biodiversity, and may open up new options for non-human primate research models.
Previous sequencing efforts have had problems with regions of the genome that are rich in GC bases. These regions are often thought to have regulatory functions. This work has corrected many of the problems with sequences covering these GC-rich regulatory regions.
The Genome 10K Community of Scientists (G10K) was established in 2008 when genomic sequencing got cheaper and technologies got better. The aim is to eventually generate high-quality reference genomes for the known living vertebrates, about 70,000 animals. There are hundreds of scientists from over 50 institutions in a dozen countries that have been a part of the G10K-VGP consortium.
The genomic sequences have been reported in a series of papers, some in a special issue of Nature. The work was done by combining sequencing approaches that analyze long stretches of nucleotide bases. In the early days of next-generation sequencing, which sped up the process and made whole-genome sequencing possible, genomes were digested into small bits that were sequenced and then pieced together. with computational tools. While that approach was efficient, it made reading repetitive, long sequences challenging or even impossible. Now, long-read methods have filled in many of the gaps in these repetitive stretches.
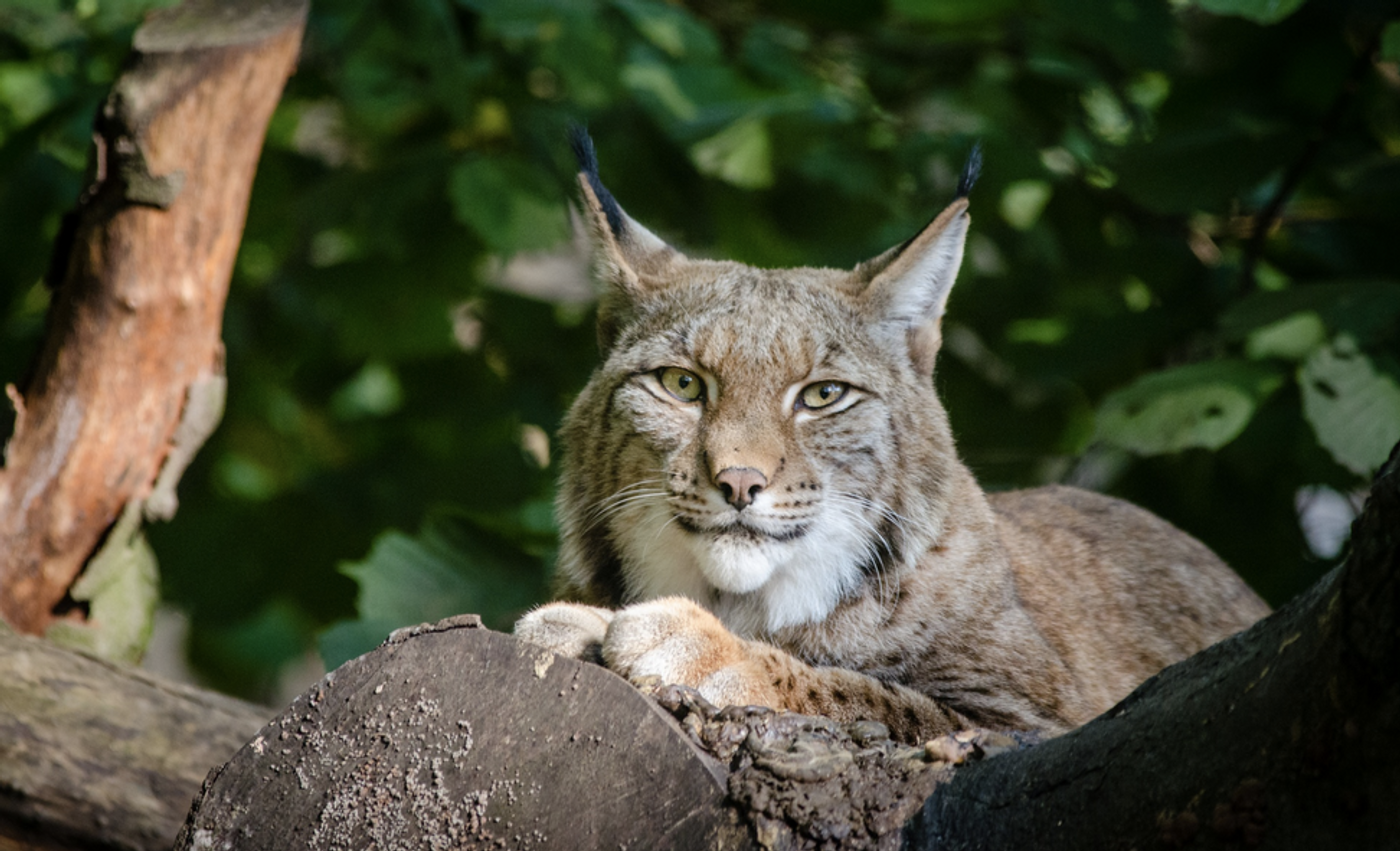
Some animals that were included in this work are the Zebra finch, Anna's hummingbird, Egyptian fruit bat, Canada lynx, vaquita, platypus, and the Zig-zag eel, among others.
Phase 1 of the project is seeking to sequence one representative from each of the 260 vertebrate orders. Phase 2 will eventually focus on getting representative species from each family.
"Our new approach to produce structurally validated, chromosome-level genome assemblies at scale will be the foundation of ground-breaking insights in comparative and evolutionary genomics," said Kerstin Howe, lead of the curation team at the Wellcome Sanger Institute in the UK.
The work has also cleaned a lot of false duplications out of reference genomes. "We have thousands of genes in the literature that are false duplications. The genes are not actually there!" said Rockefeller University's Erich D. Jarvis, Chair of the Vertebrate Genomes Project. "It is unconscionable to be working with some of these genomes."
"Completing the first vertebrate reference genome, human, took over ten years and three billion dollars. Thanks to continued research and investment in DNA sequencing technology over the past 20 years, we can now repeat this amazing feat multiple times per day for just a few thousand dollars per genome," noted Adam Phillippy, chair of the VGP genome assembly and informatics working group and head of the Genome Informatics Section of the National Human Genome Research Institute, NIH.
Sources: AAAS/Eurekalert! via Nova Southeastern University, Rockefeller University, Nature