Expansive RNA Atlas Includes Coding & Non-Coding Molecules
We've sequenced the human genome, even the parts that are highly repetitive, don't code for protein, and are extremely challenging to analyze. Now it's time for scientists to investigate the parts of the genome that are expressed as RNA molecules, called the transcriptome, which can have a variety of different functions. What makes this especially difficult is that the transcriptome can be very different from one cell type to another, and can even vary at the single-cell level.
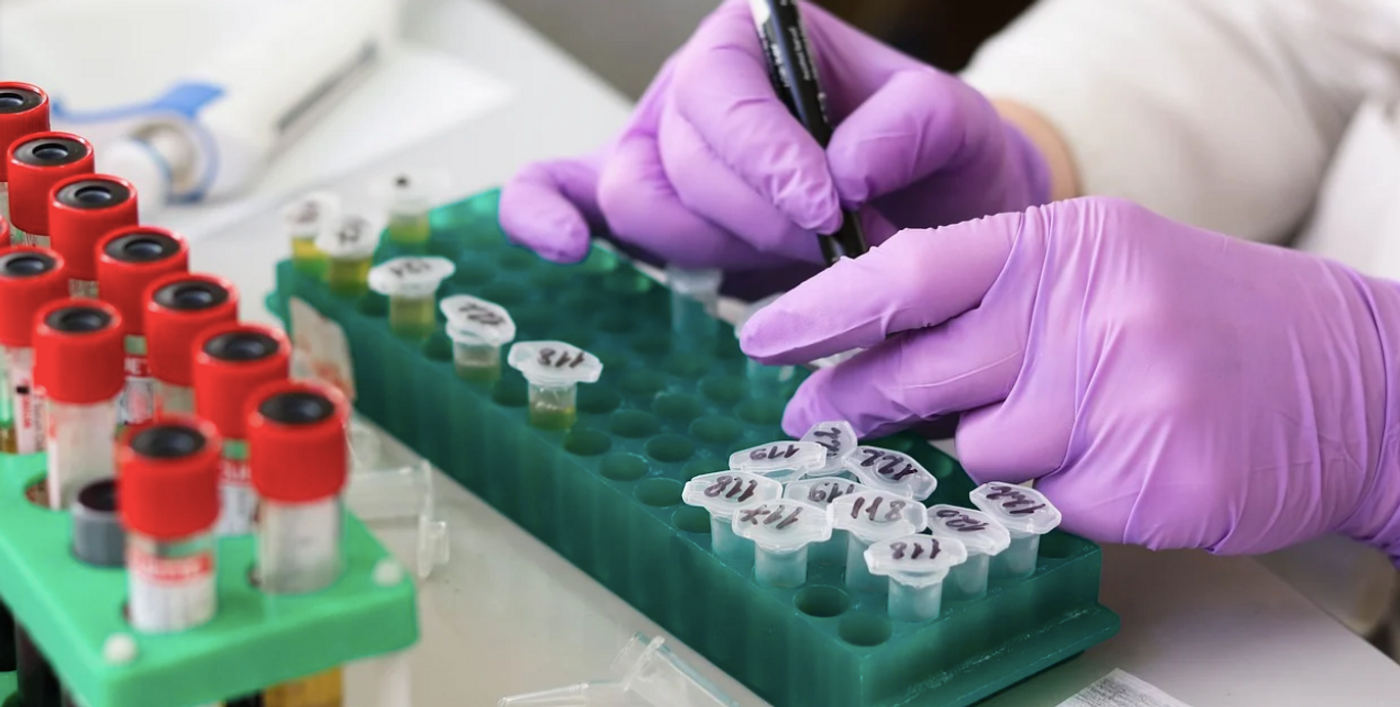
Scientists are making serious efforts to create transcriptome atlases. After years of work, researchers have now made an RNA atlas using human samples, which includes not only the sequences that code for protein, but also non-coding RNA molecules. Just like coding sequences, the non-coding sequences can be different lengths and make molecules that are differently shaped; there are circular, linear, short, and long non-coding RNAs. This work, reported in Nature Biotechnology, has assessed RNA molecules from 300 different types of human cells and tissues, with data from three sequencing techniques.
"There have been other projects to catalog our transcriptome but the RNA-Atlas project is unique because of the applied sequencing methods," said Professor Pieter Mestdagh from the Center for Medical Genetics at Ghent University. "Not only did we look at the transcriptome of as many as 300 human cell and tissue types, but most importantly, we did so with three complementary sequencing technologies, one aimed at small RNAs, one aimed at polyadenylated (polyA) RNAs, and a technique called total RNA sequencing."
Total RNA sequencing, which includes many types of RNA molecules from a cell, like ribosomal RNA and precursor RNA, enabled scientists to reveal the existence of non-coding RNAs, some of which have important regulatory functions. There are variations in ncRNAs that have been linked to some diseases.
By using three sequencing methodologies to compare data, the researchers could get an idea of the relative levels of the ncRNAs. They were also able to find some characteristics of these ncRNAs too, like whether they are circular or linear or capped with a poly-A tail. In some cases, they learned more about the function of the RNA molecules; the relative abundance may indicate which are regulatory molecules and whether they have an influence during the transcription of DNA, or during the processing of RNA.
The work is available on the R2 web portal.
"By combining all data in one comprehensive catalog, we have created a new valuable resource for biomedical scientists around the world studying disease processes. A better understanding of the complexity of the transcriptome is indeed essential to better understand disease processes and uncover novel genes that may serve as therapeutic targets or biomarkers," said Professor Pavel Sumazin of the Baylor College of Medicine.
"The age of RNA therapeutics is swiftly rising. We've all witnessed the impressive creation of RNA vaccines, and already the first medicines that target RNA are used in the clinic. I'm sure we'll see lots more of these therapies in the next years and decades."
Sources: AAAS/Eurekalert! via Ghent University, Nature Biotechnology