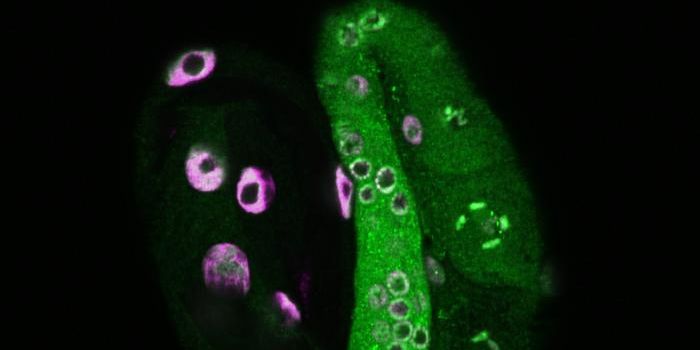
DNA, the diverse combination of just four nucleotide letters that writes our genetic code, can do more than just form a double helix. In a new study from the National Institutes of Health (NIH) and the National Center for Biological Sciences (NCBS), scientists look at the pattern of gene expression that could potentially be impacted by various supercoiling arrangements.
To fit into tight-knit cells, DNA has to be packed in to the small space. However, during events like regular DNA replication and reproduction, the genetic material has to be unpacked and unwound in order for other compounds to be able to read and create RNA and proteins from DNA.

In the NIH/NCBS study published in the journal
Nature Communications, the authors described a “constant state of structural flux” that goes hand in hand with the state of cellular activity, indicating that the genetic code can respond to changes in order to improve body-wide function.
Studies in human and yeast cells investigating the supercoiling of DNA have been completed, but the present study looks at the genome of a favorite model organism,
Escherichia coli.
In addition to being a commonly used model organism,
E. coli cause bacterial infections, often from contaminated food or drink, leading to diarrhea, urinary tract infections, pneumonia, and more (
Centers for Disease Control and Prevention).
Bacteria experience environmental influences just like humans do, and the current study is considering the possibility that by altering their gene expression, bacteria could increase their chance of survival despite any environmental hardships like starvation, lack of oxygen, or unusual temperatures. Scientists believe that there is a supercoiling connection to bacterial sensitivity to changes in environmental influences.
The method presented in the study started with exposing
E. coli genetic material to a chemical called trimethylpsoralem, whose bondage to DNA is a representation of how much DNA supercoiling is occurring. Next comes ultraviolet light and microarray technology, focused on two different populations of
E. coli bacteria so the researchers could visualize a distinction between the activity of DNA under two opposite environmental conditions.
After securing a fine-scale resolution view of supercoiling patterns, researchers soon gained insight into section-specific genomic information.
One group of bacteria were cultured in a nutrient-rich environment, one which promoted active division and a continuously growing population. The other group struggled to grow in a nutrient-poor environment and existed as a stationary population as opposed to actively growing.
For the health, actively dividing group of bacteria, the entire genome appeared supercoiled, while the nutrient-deprived group showed a diverse gradient of supercoiling.
This study provides proof-of-concept that the supercoiling of a genome is not uniform and that it varies locally across genes,” said first author of the study, Avantika Lal, PhD. “It also provides evidence to support the hypothesis that bacterial cells could be regulating gene expression and their own physiologies by altering the structure of their genomes."
Source:
National Center for Biological Sciences