One of the first research Institutes to be part of the MinION Access Programme (MAP), The Genome Analysis Centre (TGAC), UK, plans to use the miniaturized sequencing device to conduct live environmental surveillance--rather than gathering samples to take back to the laboratory--enabling the researchers to deliver real-time experimental genetic data for immediate analysis.
MAP is a community-focused access project started in Spring 2014 that aims to enable a broad range of people to explore how the MinION may be useful to them, to contribute to developments in analytical tools and applications, and to share their experiences and collaborate.
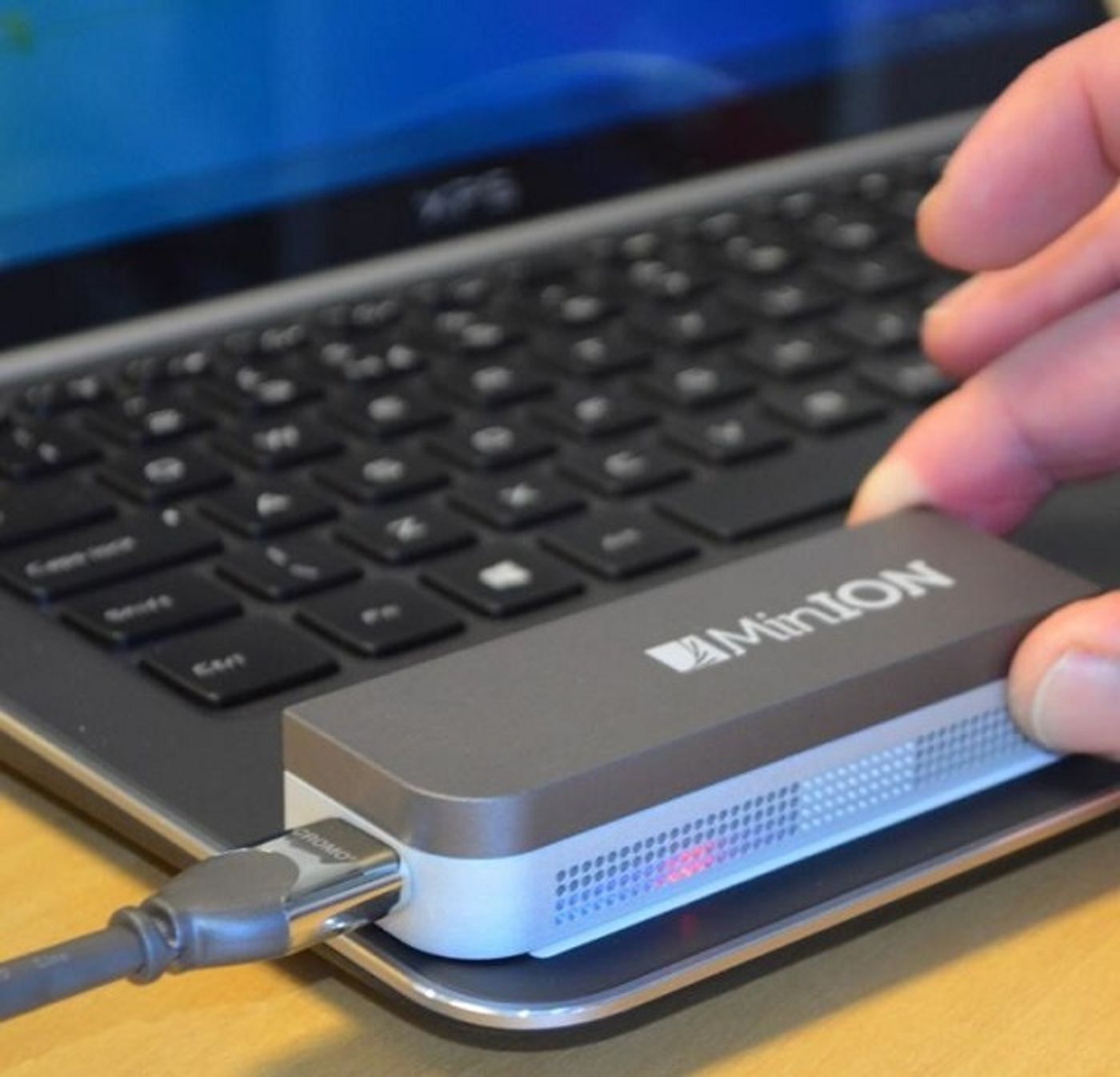

The MAP is so far focused on DNA sequencing, and in time is expected to encompass the direct analysis of RNA and proteins. Listening to this community helps Oxford Nanopore, UK, a company that's developing a proprietary technology platform for the direct, electronic analysis of single molecules, to continuously improve its products.
The team of scientists from TGAC's Data Infrastructure and Algorithms and Plant and Microbial Genomics groups trialled the miniaturized sensing system by sequencing environmental samples, containing DNA from hundreds or thousands of different organisms.
The team experimented first with a mock community, where they used a simple set of DNA samples from 20 bacteria, created for the Human Microbiome Project. Having developed their experimental and data methods, they then tested real environmental samples sequencing them on the MinION and Illumina platforms for comparison.
Their aim was to sequence the microscopic biological molecules in the air around us -- bacteria, spores, and viruses. Many key crop diseases spread via the air as spores, as well as some animal and human diseases. Analyzing such samples triggers technical issues, where there are very low levels of biological material present when sequencing DNA from air.
Although the scientists faced challenges working with complex metagenomic (genetic material recovered directly from an environment) samples in live-time, the introduction of the MinION as a potential portable laboratory made a major impact to the research's goal.
"In-field surveillance presents a number of hurdles. With its compact size, cheap device cost, simple library preparation and streaming nature, the MinION provides a significant step towards addressing these challenges," says Richard Leggett, PhD, member of the MAP task force.
Leggett and the rest of the task force recently presented the TGAC team's preliminary findings at the Advances in Genome Biology & Technology conference in Florida, in a talk titled "Towards Real-Time Surveillance Approaches Using Nanopore Sequencers."
"Our research is still at an early stage, but indications are that the MinION is capable of sequencing complex metagenomic samples," Leggett says. "It's great to be part of the MinION Access Programme, as it has provided us with access to the very latest sequencing technology. Though the MinION is not perfect, current sequencing platforms are just too large and cost too much to purchase to consider deploying in-field."
Having the ability to deploy real-time monitoring in the field could change human disease epidemics, he notes. "These are early days, but it looks like a combination of a small, low-power sequencing device with the correct experimental and computational approaches can spot pathogens," says Matt Clark, head of the MAP task force and Plant and Microbial Genomics Group Leader at TGAC.
In addition to studying metagenomics, the TGAC MAP team are also interested in understanding new technologies such as the MinION and providing the scientific community with tools to help them get the most out of these technologies.
[Source: TGAC]