Rapid Idenditfication of Microorganisms in Positive Blood Cultures Using MALDI-TOF MS
Conventional methods of identifying a positive blood culture require waiting for the blood culture broth to grow the organism and once positive, it has to be subcultured to specific media such as blood, chocolate, and MacConkey's agar. A Gram stain performed on the broth gives only basic information--Gram positive or Gram negative cocci or rods and yeast or fungi. Fast-growing organisms including most of the Enterobacteriaceae and Staphylococcus aureus can be detected in 8-12 hours. However, after subculturing the broth, the plates have to be incubated 18-24 hours before identification and antibiotic susceptibilities can be performed, which then take another 18-24 hours. The length of time from specimen collection to final identification with susceptibility patterns takes up to 48 hours for organisms that grow quickly. Many bacteria can take days to grow delaying identification even longer.
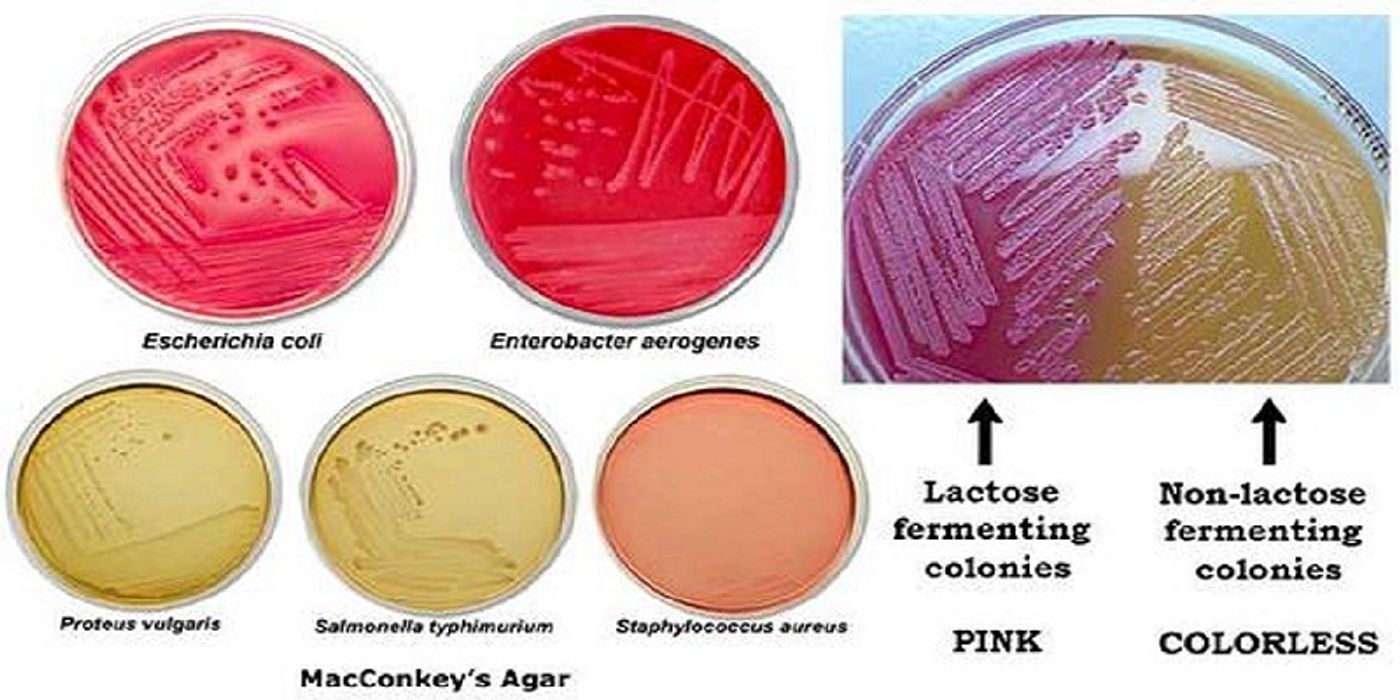
Figure 1. Gram Negative Rods on MacConkey's Agar
Sepsis is a leading cause of death in both children and adults with the mortality rate as high as 50%. Early recognition of sepsis and administration of antibiotics are crucial to mitigate mortality rates. The first 24 hours is the most critical period to identify the presence of sepsis and begin treatment with appropriate antibiotics. It's estimated that for every hour antibiotics are delayed, survival decreases by 7.6%, and if the wrong antibiotic is used, there is a fivefold reduction in survival. Most patients are started on a broad-spectrum antibiotic, but because of increasing antibiotic resistance, these may be ineffective. The need for rapid turnaround time for identification and susceptibility testing has the potential to increase survival rates.
Matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF-MS) has emerged within the last decade as a powerful tool for rapid and accurate identification of microorganisms. The process of microbe identification using MALDI-TOF-MS technology is a complex process. Singhal et al.describe the analytic steps as follows:
"The sample for analysis by MALDI MS is prepared by mixing or coating with solution of an energy-absorbent, organic compound called matrix. When the matrix crystallizes on drying, the sample entrapped within the matrix also co-crystallizes. The sample within the matrix is ionized in an automated mode with a laser beam. Desorption and ionization with the laser beam generates singly protonated ions from analytes in the sample. The protonated ions are then accelerated at a fixed potential, where these separate from each other on the basis of their mass-to-charge ratio (m/z). The charged analytes are then detected and measured using."
Although the initial investment in the equipment needed to perform MALDI-TOF-MS is high, the cost-effectiveness is realized from its rapid and accurate identification methods.
While the use of samples for MALDI-TOF-MS identification taken from bacterial colonies grown on plates is well-documented in the literature, a method to test positive blood cultures directly from the broth is needed to reduce the time from blood collection to antibiotic administration significantly. Bazzi et al. analyzed the accuracy of their proposed "direct method of lysis centrifugation" when compared to the gold standard of testing from a colony that has grown in culture overnight. Samples included 60 Gram-positive and 60 Gram-negative isolates that were analyzed with the Vitek MALDI-TOF-MS.
Their proposed method involved "centrifugation followed by washing and 30% acetic acid treatment along with our modified lysis centrifugation followed by washing and 30% formic acid treatment method, lysis centrifugation followed by washing and 100% ethanol treatment method, and picking colonies from 90 to 180 min subculture plates in order to determine the accuracy and precision of each method."
Their data revealed that the methods of (1) extraction by lysis centrifugation followed by washing and 30% formic acid treatment method or (2) 30% acetic acid treatment were both accurate for a range of Gram-positive and Gram-negative bacteria. If microbiology laboratories adopt these methods, the cost of treating sepsis will be reduced, and its overall impact on survival rates will increase.