A New Way to Detect Any Human Virus
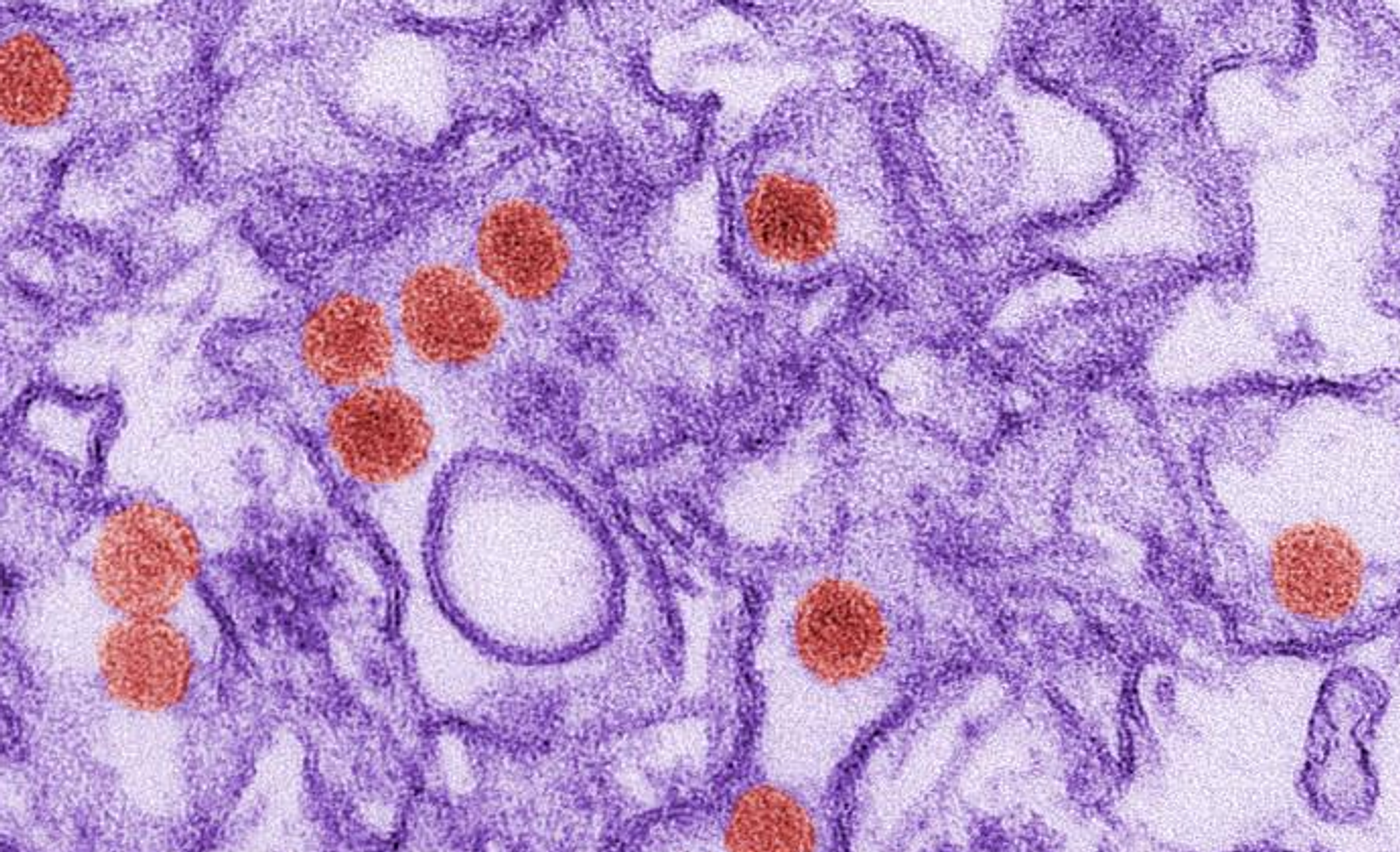
Some viruses, like Zika, can be challenging to study when they suddenly cause outbreaks because they can be tough to detect in blood samples. Now, scientists in the lab of Pardis Sabeti at there Broad Institute have created a way to design molecular baits that can grab viral particles. The new tool, called CATCH, will enable researchers to study viruses that are only present at low levels in clinical samples. The work has been reported in Nature Biotechnology. Sabeti discussess how we monitor and respond to infectious disease outbreaks in the video.
"As genomic sequencing becomes a critical part of disease surveillance, tools like CATCH will help us and others detect outbreaks earlier and generate more data on pathogens that can be shared with the wider scientific and medical research communities," said the co-senior author of the study Christian Matranga, who now works with a biotech startup.
Viral particles can be captured, even at low levels, with metagenomic techniques, which aim to sequence all of the genetic material in a sample. But those low-level viruses can get lost in the huge amount of genetic information from other sources, like microbes and patient DNA.
Scientists in the Sabeti lab needed a way to use probes they had already made for detecting pieces of viral DNA, while also enriching the microbes that were present at low levels, like Zika.
"We wanted to rethink how we were actually designing the probes to do capture," said MIT graduate student Hayden Metsky. "We realized that we could capture viruses, including their known diversity, with fewer probes than we'd used before. To make this an effective tool for surveillance, we then decided to try targeting about 20 viruses at a time, and we eventually scaled up to the 356 viral species known to infect humans."
They developed CATCH (Compact Aggregation of Targets for Comprehensive Hybridization), which enables a user to design probes that grab genetic material from microbial species, including viruses that infect people. The probes are based on any human virus that's in the GenBank database (from the National Center for Biotechnology Information) and can be made by any number of commercial probe-synthesis companies.
In this work, the researchers used CATCH to design probes for Zika and chikungunya and were able to detect the presence of Zika viruses in several regions before other scientists were able to. This demonstrates that this strategy could help control outbreaks in the future.
Ideally, this approach would be used in locations where viral outbreaks are common. "We'd like our partners in Nigeria to be able to efficiently perform metagenomic sequencing from diverse samples, and CATCH helps them boost the sensitivity for these pathogens," explained postdoctoral researcher Katie Siddle.
The method can also help clinicians who encounter fevers that may be caused by a virus, but are difficult to diagnose. "We're excited about the potential to use metagenomic sequencing to shed light on those cases and, in particular, the possibility of doing so locally in affected countries," said Siddle.
CATCH can also easily be used when new viral sequences are discovered. The probes designed by Metsky and Siddle have also been made available for public use.
This work may not only improve our ability to watch for viral outbreaks, but it could also aid in large-scale studies of microbiomes or become a diagnostic tool.
Sources: AAAS/Eurekalert! Via Broad Institute of MIT and Harvard, Nature Biotechnology