Scientists Capture Video of a Virus Forming
In a first, scientists have captured video of individual viruses as they form, illustrating viral assembly. This work can help us learn more about how we can combat viruses, and how to engineer nanoparticles that can assemble themselves. This work focused on simple single-stranded viruses that are made of RNA, which are the most common kind of virus that we know of; these viruses cause the common cold, West Nile, gastroenteritis, and polio, among others infections. The findings have been reported in the Proceedings of the National Academy of Sciences (PNAS).
"Structural biology has been able to resolve the structure of viruses with amazing resolution, down to every atom in every protein," said Vinothan Manoharan, the Wagner Family Professor of Chemical Engineering and Professor of Physics at the Harvard John A. Paulson School of Engineering and Applied Sciences. "But we still didn't know how that structure assembles itself. Our technique gives the first window into how viruses assemble and reveals the kinetics and pathways in quantitative detail."
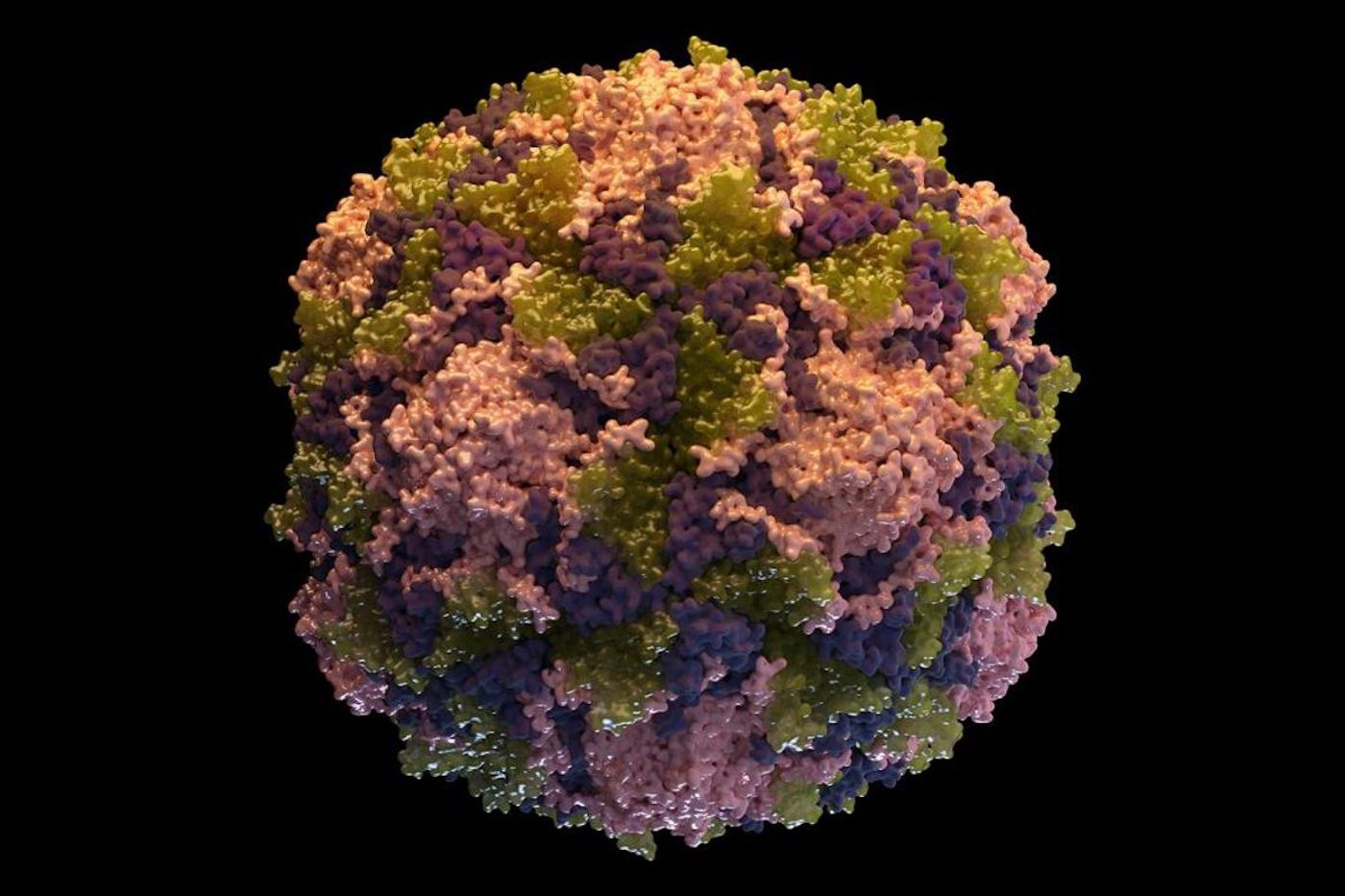
RNA viruses are typically simple structures. Manoharan’s team focused on one that can infect a bacterium. It’s a single strand of RNA that’s around 30 nanometers in diameter, and it can generate 180 identical proteins. These proteins can arrange themselves into pentagons and hexagons, forming a capsid - a protective structure that encases the viral RNA.
Until now, no one has visualized the formation of the structure. Viruses have tiny components and have been challenging to observe in real-time. Scientists turned to a tool called interferometric scattering microscopy; in it, light scatters off an object and makes a dark spot against a bright field. The physical structure of the virus can’t be seen directly, but the changes in its size can be visualized over time, as shown in the video.
In this work, strands of virus were attached to a substrate while the scientists moved proteins over the surface. With the interferometric microscope, they could watch as dark spots arose and grew darker until they reached the size of a fully-formed virus. The intensity of the darkness could be measured, enabling the scientists to calculate the number of proteins attaching to the strands of viral RNA over time.
"One thing we noticed immediately is that the intensity of all the spots started low and then shot up to the intensity of a full virus," Manoharan noted. "That shooting up happened at different times. Some capsids assembled in under a minute, some took two or three, and some took more than five. But once they started assembling, they didn't backtrack. They grew and grew, and then they were done."
Their results were compared to two other simulations with different predictions. The first one was ruled out, but the second predicted pathway agreed with their observations; the proteins form a critical mass called a nucleus before the capsid can form. Once the nucleus is generated, which happens at different times with different viruses, the growth of the virus continues until the correct size is reached. The scientists also noticed that if more proteins are moving on the substrate, the assembly can go awry.
"Viruses that assemble in this way have to balance the formation of nuclei with the growth of the capsid. If nuclei form too quickly, complete capsids can't grow. That observation might give us some insights into how to derail the assembly of pathogenic viruses," Manoharan explained.
Now that the assembly pathway has been revealed, scientists can explore more questions and learn more about how to model the mechanisms. This work could also be useful in the design of nanomaterials.
"This is a good example of quantitative biology, in that we have experimental results that can be described by a mathematical model," said Manoharan.
Learn more about viruses from the video.
Sources: AAAS/Eurekalert! via Harvard John A. Paulson School of Engineering and Applied Sciences, PNAS