The Microbial Communities That Form on the Tongue
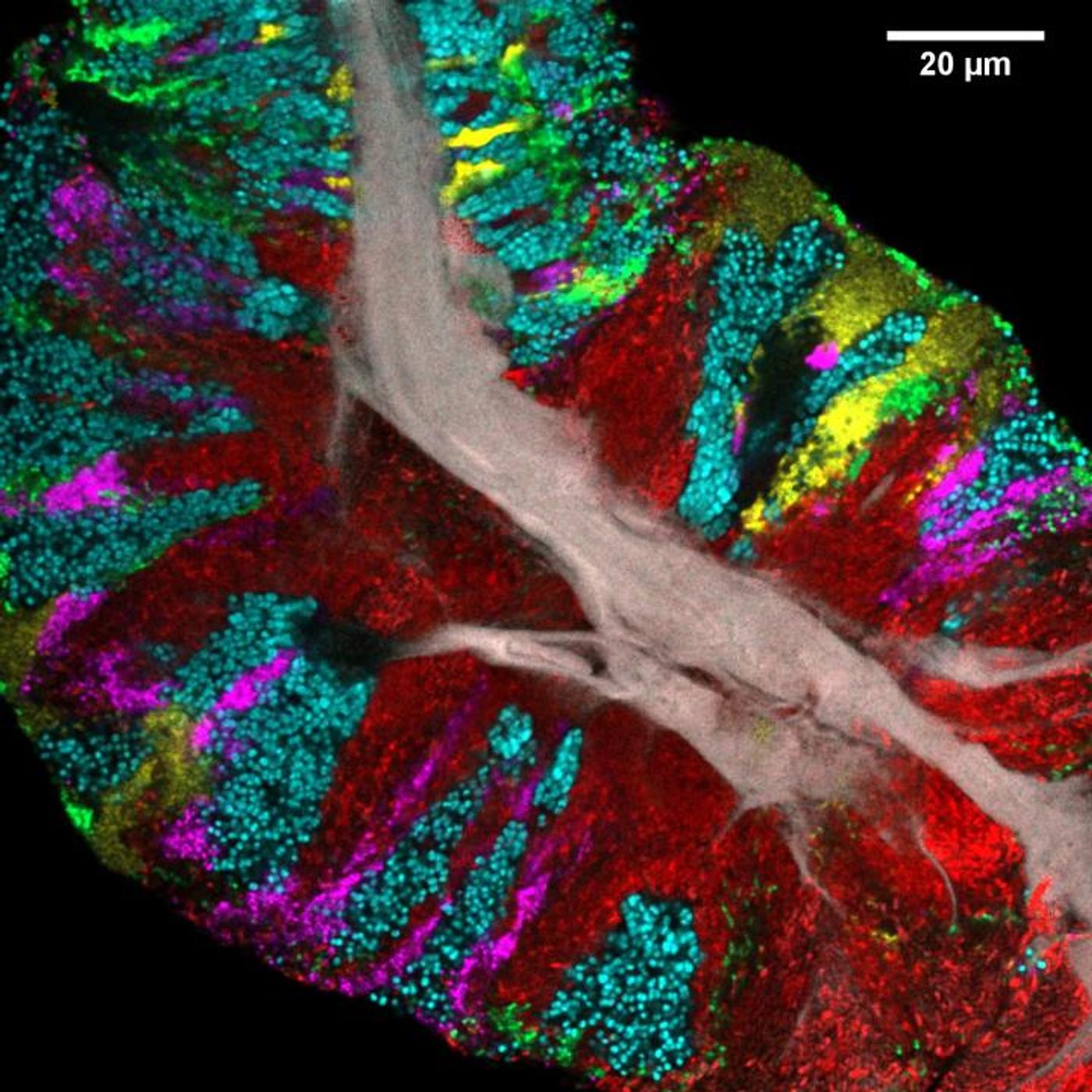
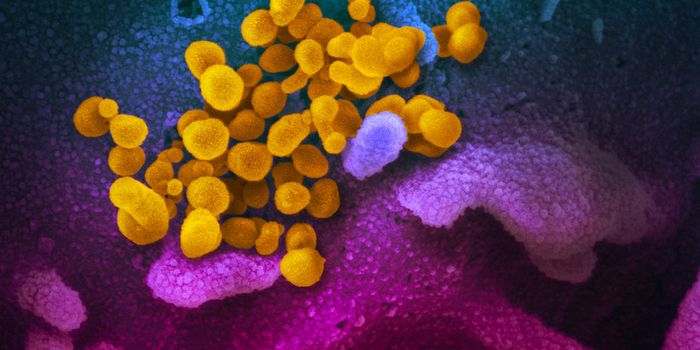
Scientists used a fluorescent imaging tool to analyze how bacteria grow on the human tongue and found that these microbial communities have a very structured organization. The human mouth hosts a complex world of bacterial biofilms that are affected by a huge number of factors including oral hygiene, moisture, temperature, oxygen levels, and disruptions like abrasions. The findings have been published in Cell Reports.
"From detailed analysis of the structure, we can make inferences about the principles of community growth and organization," said the senior study author Gary Borisy, of the Forsyth Institute and the Harvard School of Dental Medicine. "Bacteria on the tongue are a lot more than just a random pile. They are more like an organ of our bodies."
The microbes in the oral microbiome impact one another. They all carry their own genes and can metabolize molecules they encounter, secrete compounds they produce, and physically interact with one another. Some microbes may compete for certain binding spots on the tongue, but the patterns that emerge have not been well-studied.
"We think that learning who is next to who will help us understand how these communities work," said study co-author Jessica Mark Welch, a microbial ecologist at the Marine Biological Laboratory in Woods Hole, Massachusetts. "The tongue is particularly important because it harbors a large reservoir of microbes and is a traditional reference point in medicine. 'Stick out your tongue' is one of the first things a doctor says."
In this work, the scientists utilized Combinatorial Labeling and Spectral Imaging - Fluorescence in situ Hybridization (CLASI-FISH), a recently developed technique from the Borisy lab, in which microbes are labeled with fluorophores to enable the simultaneous identification and localization of different microbes.
"Our study is novel because no one before has been able to look at the biofilm on the tongue in a way that distinguishes all the different bacteria, so that we can see how they arrange themselves," Borisy explained. "Most of the previous work on bacterial communities used DNA sequencing-based approaches, but to get the DNA sequence, you have to first grind up the sample and extract the DNA, which destroys all the beautiful spatial structure that was there. Imaging with our CLASI-FISH technique lets us preserve the spatial structure and identify the bacteria at the same time."
Researchers took tongue scrapings from 21 healthy volunteers, and genetic sequencing data was used to identify the bacteria present in the samples. There were 17 bacterial genera that were commonly found as either free-floating bacteria, microbes bounds to host cells, or biofilms; they were identified in over 80 percent of study participants. All study volunteers carried three genera: Actinomyces, which was usually near the core, Rothia, that tended toward the exterior and Streptococcus, which tended to form interior patches or a thin crust on the exterior.
"Collectively, our species-level imaging results confirm and deepen our understanding of habitat specificity of key players and show the value of investigating microbiomes at high imaging and identification resolution," Mark Welch said.
This work suggests that individual bacterial cells or small clusters or them bind to the tongues epithelium. These cells proliferate depending on how they interact and the microenvironment that is present, creating a patchwork of larger groups.
Get a look at some of the many microbes that can be found in the human mouth in the video above.
Sources: AAAS/Eurekalert! via Cell Press, Cell Reports