Modeling the Evolution of Antibiotic Resistance
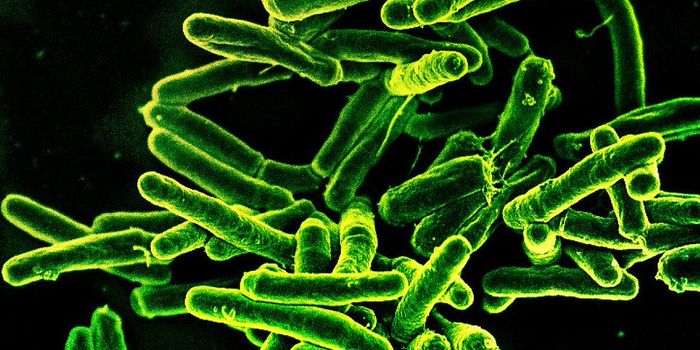
Pathogenic bacteria are becoming increasingly resistant to standard antibiotics, for a variety of reasons. Scientists are trying to learn more about how microbes gain the ability to evade the effects of these important drugs. In one recent study reported in Nature Communications, scientists subjected colonies of Escherichia coli bacteria to 95 different antibiotics. They used robotic technologies to grow them for 250 generations and analyzed how gene expression changed in the microbes over time.
"Laboratory evolution combined with genomic analyses is a promising approach for understanding antibiotic resistance dynamics," explained study leader Tomoya Maeda of RIKEN BDR.
They developed gene expression profiles for the microbes and used computational tools to analyze this vast amount of data.
"We found that E. coli's evolutionary dynamics is attributable to a relatively small number of intracellular states, indicating that it is likely equipped with only a limited number of strategies for antibiotic resistance," said Maeda. If the researchers can quantify how antibiotic resistance evolves in E. coli, the team is hopeful that they will be able to predict and thereby control antibiotic resistance.
The scientists were able to use their system to test 2,162 drug pairs, and this identified 157 pairs that could help suppress the acquisition of antibiotic resistance in E. coli.
"We believe that our results can be applied to the development of alternative strategies for suppressing the emergence of drug-resistant bacteria," said Maeda.
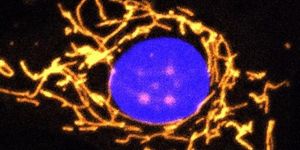
In another study reported in RSC Chemical Biology, scientists engineered fluorescent probes that can mark bacteria, and as such can be used to observe how bacteria react to antibiotics. This effort is meant to get a better look at the behavior of individual microbes rather than large colonies.
“By using an antibiotic tagged with a fluorescent probe, we could see the rate at which the antibiotic was being taken up by the bacteria. While one group of bacteria started glowing at the same time, indicating they were taking up the bacteria simultaneously, another group was very variable as to when or if they picked up any of the probes,” noted study author Dr. Mark Blaskovich of the University of Queensland. "This variability might explain why these bacteria are able to survive – they aren’t picking up any of the antibiotics at all.”
More work will be needed to learn why some bacteria are unaffected by exposure to antibiotics. Such research could help us learn more about improving antibiotic efficacy.
Sources: AAAS/Eurekalert! Via RIKEN, University of Queensland, RSC Chemical Biology, Nature Communications