Decontaminating the Cancer Microbiome
Valid scientific conclusions require a solid foundation of good data that comes from reliable techniques. Studies that investigate microbial genetics have to ensure that the microbes that fill our world and our bodies don’t contaminate the samples that are under study. Researchers that are trying to learn more about the microbial communities that reside in cancerous tumors have now created a system for eliminating contamination in microbial genetic data contained in The Cancer Genome Atlas (TCGA). They are trying to get a clear picture of the tumor microbiome, which could improve our ability to treat these tumors or understand how they form.
After decontaminating their dataset, the researchers found that there is a slightly different group of microorganisms in tissues that are healthy compared to cancer tissues. They also found that bacteria from these unhealthy sites can migrate away into the bloodstream. The findings have been reported in Cell Host & Microbe.
"All microbiota studies are plagued by the notion that if you find a microbe, was it really in the tissue or was it contamination introduced during processing?" said Xiling Shen, the Hawkins Family Associate Professor of Biomedical Engineering at Duke University. "We've invented a method that can extract the microbes that were truly in each sample and used it to build what we've called The Cancer Microbiome Atlas, which will be a tremendous resource for the community and allow us to understand how cancer alters an organ's microbiome."
TCGA contains genomic sequencing, epigenetic, and gene expression data from more than 20,000 primary cancers that represent 33 types.
Researchers may one day be able to assess microbes in the blood to diagnose disease and predict outcomes.
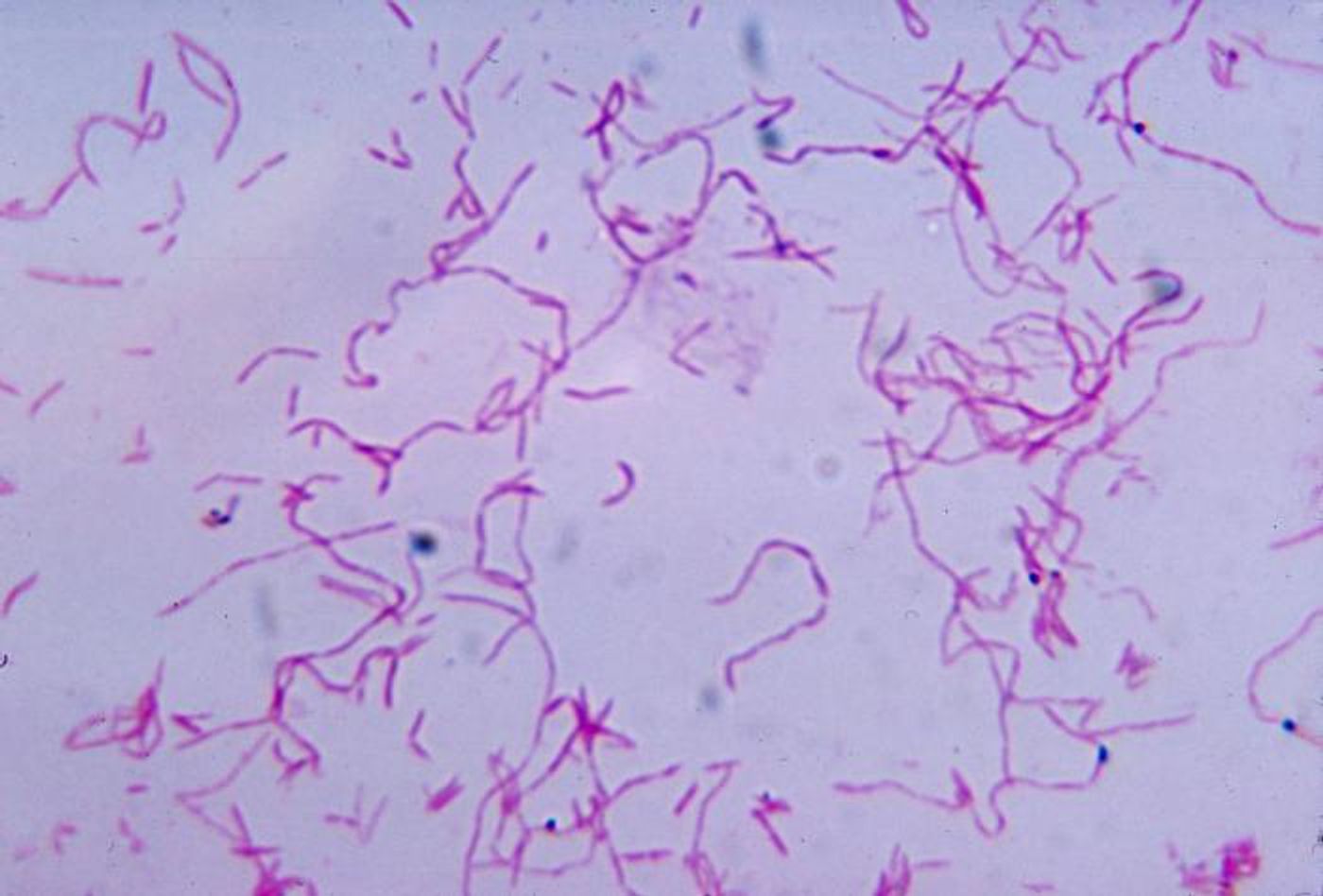
For example, researchers have shown that the level of Fusobacterium nucleatum bacteria in colorectal cancer can indicate which stage the cancer is in, whether it will metastasize, and the potential prognosis. While other studies have searched for these types of microbial biomarkers, the tiny sample sizes increase the chances for contamination, which has made it challenging to find them; microbial DNA identified in tumor samples can sometimes come from the laboratories doing the testing.
Graduate student Anders Dohlman in the Shen lab was able to compare the microbiomes from cancer tissues to those from blood. Species that appeared indiscriminately were ruled out. Then he compared data obtained at different institutions, like Harvard and Baylor Universities. Microbial species only found at one site were likely to be contaminants, he concluded. Using that, he created a contamination signature for the institutions.
"A big challenge in this process was mixed-evidence species, which are bacteria that are both a contaminant and endogenous to the tissue," said Dohlman. "But because TCGA has so many different types of data, we were able to tease it out. Big data really helps!"
After decontaminating the data, the researchers found a combination of two microbes that are unique but often found together in colorectal cancer patients. One of the species seems to be linked to patient survival. Some tumors also appear to change the microbiome of the organ in which they reside.
"There has been a sort of crisis in the field about whether or not high-profile papers can be reproduced, owing to the challenge of contamination," said Shen. "For example, while one center would be able to reproduce its results, another center would not. This explains why: Each center has its own very consistent bias. (Its own resident microbe contaminants.) In the future, new studies can use our method to remove this bias and reproduce results, and research centers might be able to use their bias we've identified to mitigate their contamination."
Sources: AAAS/Eurekalert! via Duke University, Cell Host & Microbe