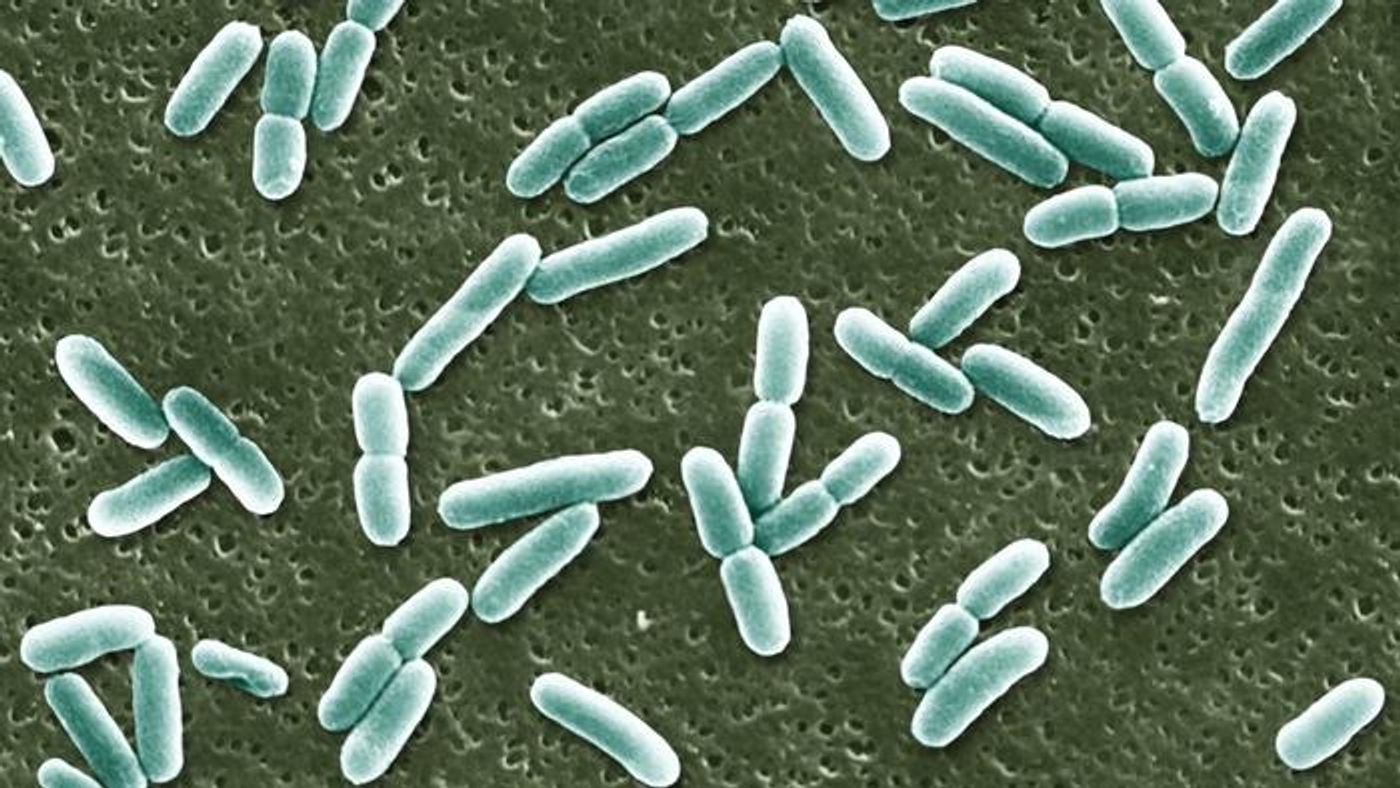
Ever wish you could predict the future? Researchers at UC Davis designed a computer model to do just that - if you’re E. coli, that is.
The model allows researchers to predict how E. coli’s metabolic pathways will respond to various experimental conditions. Think of it as a virtual science experiment. “We are exploring a vast space here”, says study author Ilias Tagkopoulos. “Our aim is to create a crystal ball for the bacteria, which can help us decide what is the next experiment we should do to explore this space better.”
The work started by creating Ecomics, a meta-database for
E. coli. Ecomics houses 4,389 gene expression profiles from 649 different conditions. It took a week to compile all of these data into Ecomics, but that’s nothing compared to the 2 years it took to build the Multi-Omics Model and Analytics (MOMA) platform. The platform actually consists of four different layers - transcriptome, proteome, metabolome, fluxome, and phenome. These layers were then integrated into a single model. “The number of layers, and the amount of data involved are unprecedented”, says Tagkopoulos.
The group needed some powerful computers to wrangle all that data. They received a grant from the National Science Foundation to use Blue Waters - one of the most powerful supercomputers in the world.
It sounds impressive, but what’s the point? The new tool could be used by researchers to make a dry run of their experiments. Instead of investing the time and money to run an experiment in multiple growth conditions, Ecomics and MOMA could help identify the best growth condition for a specific experiment.
So, does it work? It sure does. The researchers pitted their model - MOMA - against two others, ME-Model and EBA, to see how well it could predict changes in gene expression in 16 knockout mutants. MOMA outperformed ME-Model by 38-280% and EBA by 47-311%.
They aren’t stopping at E. coli, however. The group has plans to build similar predictive models for Salmonella enterica and Bacillus subtilis, two species that cause foodborne illnesses. “We’re living in an amazing era at the intersection of computer science, engineering, and biology”, says Tagkopoulos. “It’s very interesting to me.”
Sources: Nature Communications,
UC Davis