Low error rate long-reads enable the identification of long variant haplotypes in metagenome assemblies
-
Derek Bickhart, PhD
Research Microbiologist/Bioinformatician, US Department of Agriculture's Dairy Forage Research CenterBIOGRAPHY
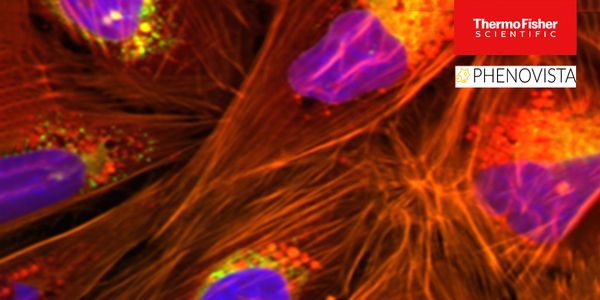
Microbial communities include distinct lineages of closely related organisms which have proved challenging to separate in metagenomic assembly. Challenges include the existence of highly related organisms that exist the same environment, or several species that exist at very low abundances and are therefore inconsistently sampled. Many of these challenges are exacerbated by limitations in DNA sequencing technologies, where some platforms produce reads that are too short or reads that have too many errors. The advent of long, accurate “HiFi” reads presents a possible means to address the challenge of metagenome assembly by disambiguating individual reads or contigs within metagenomic bins. We present a metagenomic assembly of a complex microbial community from parasite-infested sheep fecal material. We found that assemblies made from HiFi reads were able to properly assemble low abundance species in our sample, and could often assemble microbial genomes into single, circular contigs. Taking advantage of the low error rates of HiFi reads, we developed a method that can track linked single nucleotide polymorphisms (SNPs), which is termed “phasing,” and identify individual SNP haplotypes that indicate subpopulations of microbes in a sample. This method is sensitive and scalable, and was able to phase haplotypes as long as 309 SNPs across hundreds of thousands of bases in the microbial genome. We also report successful characterization of multiple, closely-related microbes within a sample with potential to improve precision in assigning mobile genetic elements to candidate host genomes within complex microbial communities.
Learning Objectives:
1. Identify the challenges of metagenome assembly
2. Quantify the benefits of low error rate long reads on metagenome sequencing and assembly
3. Understand concepts such as “phasing” and “haplotyping” as it relates to metagenome assembly
Sponsored By: Thermo Fisher Scientific
Please update your information
Certificate of Attendance
DOWNLOAD CERTIFICATE
Finish Registering
-
SEP 03, 2024Microbiology Week Virtual Event Series 2024
-
SEP 18, 2024Cell Biology Virtual Event Series 2024
-
SEP 26, 2024Olink Proteomics World
- See More
-
JUL 26, 2024
- See More