Using Deep Learning to Diagnose Breast Cancer Sub-Types
Cancer and computer programming may not sound like two things that go together. However, advanced programming techniques are now able to utilize large data training sets and “learn” how to solve problems. This is called machine learning. This technique is often used in things like search engine refinement, ad targeting, and even simple artificial intelligence programs. A group from California proposed that deep learning, a form of machine learning, could be used to diagnose the sub-clinical types of breast cancer.
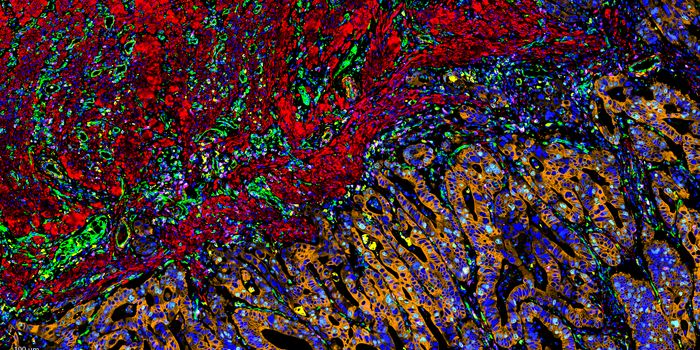
The hypothesis was that deep learning might be able to identify histological differences between estrogen, progesterone, and Her2 (ER/PR/Her2) receptor-based breast cancers. Each of these sub-types has minor characteristic differences and sometimes different treatment paths. Current diagnostic methods involve immunohistochemistry against the receptors. The deep learning program would focus on the analysis of a sample's histology using hematoxylin and eosin (H&E) images to predict the sample’s sub-clinical typing. Histological characteristics are not usually used to diagnose the sub-types as the interpretation of a sample could easily differ between doctors. The program would be able to overcome this bias.
The first step was to obtain a set of data for the deep learning program to use to train itself. Finding this dataset itself was a bit of a challenge, due to many H&E images available not being properly labeled. Initially, they ordered tissue microarrays (TMA), each with 207 samples, and put them through a set of four experiments. Experiments one and two used microarrays that had been stained by the lab, with the first experiment being only 207 samples, and the second having twenty fold more samples. The last two experiments were the same, except with the sample images colors normalized. This was done to try and make the program identify sub-types regardless of the coloration of the H&E image. The high sample normalized dataset from experiment four gave the best accuracy result at 93% correct after correcting for data bias.
Spurred on by the positive results, the group decided to test the same methodology to develop a second neural network using H&E breast cancer images from The Cancer Genome Atlas. The resulting program could predict the correct sub-clinical typing about 88% of the time. When this network was used on separate data from the Australian Breast Cancer Tissue Bank, it maintained around an 80% accuracy in its prediction of all three sub-types. They had successfully developed a new neural network that could predict the sub-type of breast cancers.
In their discussion, the team states, “Here, we illustrate a novel first step, using tissue matching to discern features that are distinctive for a patient but differ between individuals.” This implementation, once properly vetted for reliability, could aid researchers in identifying the best treatment for their breast cancer patients. It may even help pave the road for the increased use of advanced computer programs in healthcare work, which could propel healthcare into the not so far-flung future.
Sources: Nature, The Medical Futurist