Mapping the Brain
The brain still holds many mysteries, but scientists are finding ways to learn more about how it works. Researchers Catherine Dulac and Xiaowei Zhuang of Harvard have created a powerful new tool for investigating the mouse brain. Reporting in Science, they've made a cellular atlas that aims to study cellular function in its natural context. Over one million cells within a two- by two- by 0.6-millimeter block of mouse brain were examined for this research, which identified over 70 different kinds of neurons in that block and showed how they function in their location.
"This give[s] us a granular view of the cellular, molecular and functional organization of the brain - nobody had combined those three views before," said Dulac. "This work in itself is a breakthrough because we now understand several behaviors in ways that we never did before, but it's also a breakthrough because this technology can be used anywhere in the brain for any type of function."
"The problem is that people realized quite a while ago that, in order to study the brain, you need to understand its components, and those components are the cells," added Dulac, who is the Higgins Professor of Molecular and Cellular Biology and Lee and Ezpeleta Professor of Arts and Sciences. "So if you take a piece of tissue and look at the genes expressed by the cells, that tells you how many cell types there are...but that still leaves you with a big problem."
The problem is that researchers still have to remove cells from tissue samples in order to study them, she said. In doing that, valuable information, like how they are organized within that tissue, is lost.
"If you really want to understand the brain, you need the spatial context, because the brain is not like the liver or other organs, where the cells are organized in a symmetrical way," Dulac explained. "The brain is unusual in that it has this topological arrangement of neurons, so we want to be able to look at a section of the brain and see what cells are there, but also where they are and what types of cells are surrounding them."
In recent years, the Zhuang lab has engineered a tool that is perfectly applicable to this work - MERFISH - for Multiplexed Error-Robust Fluorescence In-Situ Hybridization. Zhuang had already created a super-resolution imaging tool called STORM. She wanted to see how all of the molecules work together in the cells she studied, instead of focusing on just a couple of proteins.
"We don't have just one or two different kinds of molecules in our cells, we have thousands to tens of thousands of genes that are expressed to make the molecular machinery that give cells their function," she explained. "I wanted to be able to image all those genes simultaneously, that's why we developed MERFISH."
When genes are active, they are transcribed into RNA. In MERFISH, those RNAs are tagged with barcodes. They can then be matched with a library of matching DNA probes that reveal the identities of the RNAs. The barcodes can be read through multiple imaging rounds.
"An amazing property of this method is the exponential scaling between the number of genes that can be imaged and the number of imaging rounds," Zhuang said. "If you wanted to look at 10,000 genes, you could try the brute force approach and do it one at a time, but of course no one would ever try that. The MERFISH approach is very powerful because it allows us to image and distinguish thousands of different RNAs in just about ten rounds of imaging."
Different kinds of cells express different genes, so MERFISH can identify cell types, noted Zhuang, who is the David B. Arnold, Jr. Professor of Science. MERFISH images tissue when it’s intact, so the spatial organization details are also revealed, she added. With these powerful tools, Dulac, Zhuang, and colleagues could apply them to the brain.
"There are areas in the brain that have been studied, like the cortex, where people noticed that cells were organized in a particular way, but there are a lot of brain areas for which we don't know the principles of organization," Dulac said. "The area we looked at in this study, the hypothalamus, is absolutely essential for many functions. It controls things like thirst, feeding, sleep, and social behaviors like parenting and reproduction, but we don't know how this structure is organized."
By combining MERFISH with single-cell RNA sequencing (scRNAseq), the team began to learn more. "This not only allowed the cell types to be cataloged in the hypothalamus, but also provided molecular signatures of these cell types and facilitated the selection of gene panels for MERFISH imaging," said Zhuang (who is featured in the video below).
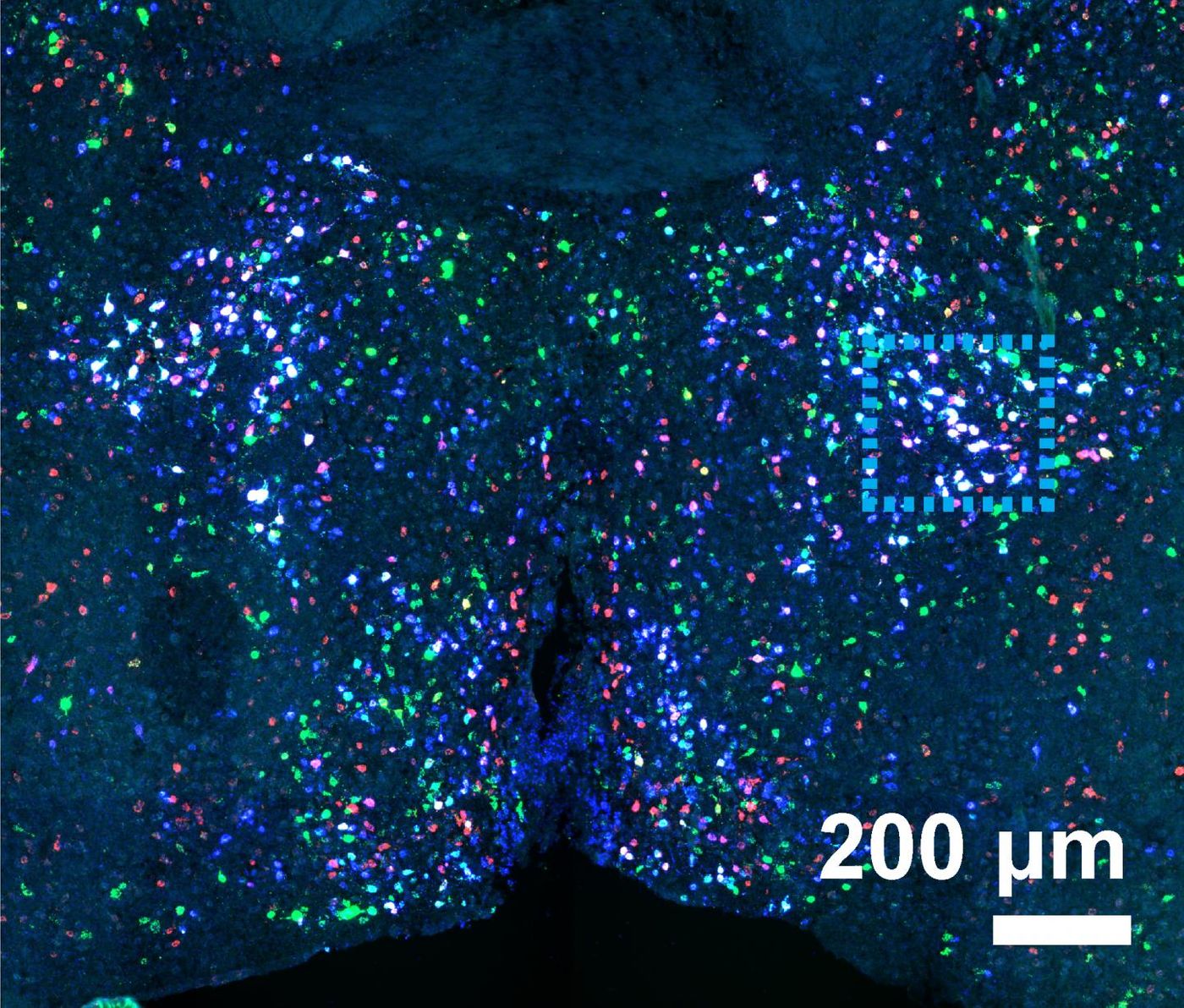
In the preoptic region of the hypothalamus, the team used MERFISH to follow over 150 genes, identifying cell types and generating a map of their locations.
"Both scRNAseq and MERFISH enabled us to identify around 70 different neuronal subtypes, most of which were previously unknown, and MERFISH imaging allowed us to additionally see the spatial distributions of all 70 neuronal types, as well as those of the non-neuronal cell types.
"What you can see is that there is an exquisite spatial organization - it leaps right out at you," Zhuang said. "You can see which neurons are neighboring each other, and not only that but because our images are molecular, you can identify how these cells are communicating with each other. Moreover, because MERFISH imaging has a very high sensitivity, we were able to identify lowly expressed genes that are critical to cell function."
One example is the c-Fos gene, which is expressed at a higher level during neural activity. Its expression indicates which neurons become active when certain behaviors start. If a research animal is allowed to behave in some way, like parenting, the cells that are active will express c-Fos, which will show which cells are involved in the parenting behavior, said Dulac. This work goes beyond that as well.
"So we can define which cells are involved in a particular behavior in ways that we could not before," Dulac said. "This is extraordinarily precise, extremely quantitative, and we can see where those cells are - so it's a cellular map, a molecular map, and a functional map, all together."
The team plans to learn more about the structure of the hypothalamus and the relationships between the cells there.
"This is really the best collaboration one could hope for," Zhuang said. "Ours are two labs whose expertise complement each other very well, and we both learned a great deal from each other. As this point, we feel like we know quite a bit about the hypothalamus, and likewise, Catherine's lab knows a great deal about MERFISH imaging, so this has been a truly exciting, rewarding process."
Sources: AAAS/Eurekalert! Via Harvard University, Science