Scientists Create Artificial Life with a Synthetic Genome
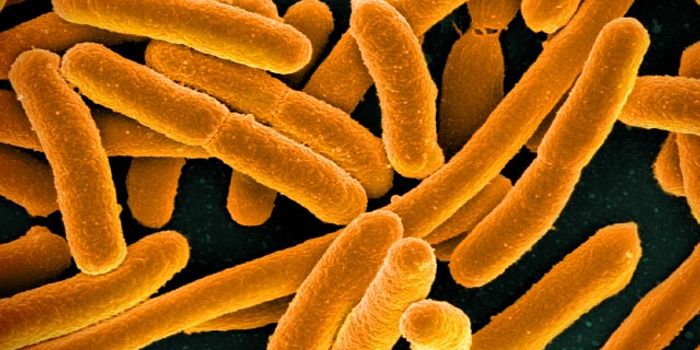
Scientists have engineered an artificial form of microbial life. Using the bacterium Escherichia coli as a model, they replaced the germ's bacterial genes with genes that had been synthesized in the laboratory. They did not stop at simply replacing the natural genome either; they edited the genome to remove redundancies. The work was reported in Nature.
In the coding portions of the genome, three nucleotide bases encode for one amino acid. Those amino acids are strung together by cellular machinery to form proteins. But there is a great deal of redundancy in the genetic code - 64 three-base-pair combinations or codons end up encoding for only 20 amino acids. That can provide some flexibility - many small errors can be tolerated, for example. But there may be other reasons.
This work, which removed those redundancies, helped reveal the potential consequences of removing them, and whether or not they are serving some other purpose. The researchers created an organism in which the genome encodes for only 61 codons, 59 of which translate to 20 amino acids, while one codon signals the start of a coding sequence and one for the stop. One transfer RNA, which moves amino acids into place, could thus be removed.
"We have stripped out some of the duplications in the natural code to make it more efficient," commented researcher Julius Fredens, who oversaw much of the research, to the BBC. In all, the team recoded 18,214 codons. They replaced the DNA in sections, eventually rebuilding the entire genome.
The engineered cell the team created is called Syn61. It doesn’t grow at the same rate as normal E. coli; it’s about 60 percent slower. The scientists don’t think that’s because of the changes in the genetic code though. Fredens suggested that they have found the small problems that are to blame, and that they can be corrected.
Since three codons are now unassigned, they could be used to introduce synthetic amino acids. Team leader Jason Chin has already done work in this area, expanding the genetic code of the nematode worm C. elegans to create proteins that can respond to light, for example.
Fredens suggested that as many as 200 synthetic chemicals could potentially be introduced into proteins. "It's pretty mind-blowing that you can expand the genetic alphabet this way," Fredens noted. "I think we're pretty far from realizing how much we can do with it, producing things we have never seen before."
Biologist Craig Venter began working on synthetic life many years ago and helped create the first synthetic cell in 2010 by transplanting a bacterial genome into a cell. In 2016, his team engineered a synthetic cell with the smallest known genome. (Learn more about Venter's work from the video above). That feat took a massive team of scientists many years, while this study was conducted by a relatively small lab using techniques that others can replicate.
Chin noted that this work should not alarm the public. "People have legitimate concerns," he acknowledged. "There is a dual use to anything we invent. But what's important is that we have a debate about what we should and shouldn't do,and that these experiments are done in a well-controlled way."