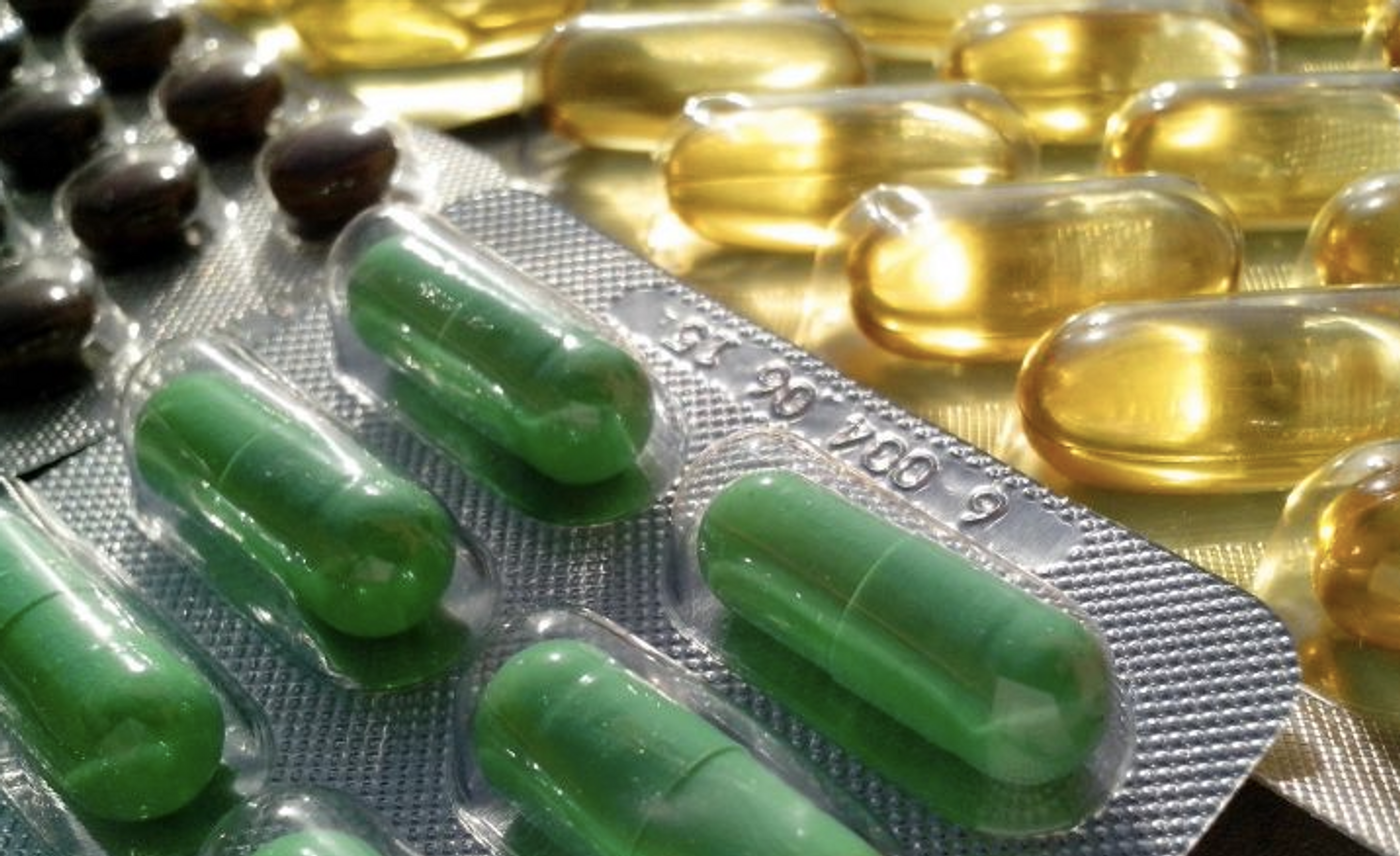
Why the Effects of a Drug Depend on Who Takes It
Many drugs can have a wide range of impacts on the patients that take them, and doctors often have to adjust a person's dosage before finding the right one. Researchers are beginning to learn more about the many reasons why this happens, like genetic or biochemical factors. Scripps Research scientists have now investigated how variations in a group of regulatory molecules called RGS proteins cause changes in the cellular response to drug exposure. The findings have been reported in Cell.
"Before you can fix things, you need to know how they're broken and how they work normally, and in this study that's essentially what we've done for these important regulatory proteins," explained study senior author Kirill Martemyanov, Ph.D., professor and chair of the Department of Neuroscience at Scripps Research's Florida campus.
There is a large family of crucial cell surface reapers called G-protein-coupled receptors (GPCRs), which are involved in many different biological functions and are associated with many disorders. Different kinds of GPCRs are activated by different substrates, like neurotransmitters or hormones. They are also common drug targets; over a third of FDA-approved medications target GPCRs.
RGS (Regulator of G-protein Signaling) proteins act like a kind of brake that reduces the action of GPCRs. Thus, GPCRs activity is restricted to a brief period so the cell has time to reset, and can then receive fresh signals. If GPCRs can't be shut down and stay on too long, downstream signaling is disrupted.
"One condition I studied earlier in my career involves the loss of RGS regulation in light-detecting cells in the retina," Martemyanov said. "Patients born with this condition can't stop perceiving light, even when they go into a dark room, and they can't track moving objects very well because they lack the normal visual refresh rate. It's easy to imagine how devastating it would be if you had a similar loss of RGS regulation in the heart or the brain where timing is so important."
While some RGS proteins have been studied, this work aimed to learn more about each one; there are twenty in human cells, which each identifies and controls a GPCR. This effort revealed a map of GPCR signaling in cells.
"This selective recognition of G-protein subunits turns out to be performed by a few elements in each RGS protein--elements organized in a pattern resembling a barcode," said the first author of the study Ikuo Masuho, Ph.D.
Phosphorylation activates GPCRs. Previous studies have suggested that patterns of phosphorylation in GPCRs create a kind of “barcode” that can be varied for specific tissues, generating targeted, physiologically relevant signals.
By analyzing the genetic data from over 100,000 individuals, the researchers found that variations and mutations in RGS barcode regions can interfere with how RGS proteins interact with GPCRs or sometimes cause them to select the wrong one. In one example, a mutation in an RGS16 barcode region that has been associated with insomnia ruins the protein's ability to recognize G proteins.
"It's clear that genetic variation in the RGS barcode regions has the potential to disrupt normal GPCR signaling, to cause disease or to create more subtle differences or traits," Martemyanov said. "For example, it may help explain why different individuals treated with the same GPCR-targeting drug often differ widely in their responses."
The work suggested that the barcode regions of RGS proteins and the receptors they regulate are continuously evolving.
Sources: AAAS/Eurekalert! via Scripps Research Institute, Cell