Extracellular Fibers Observed Moving Through the Nucleus
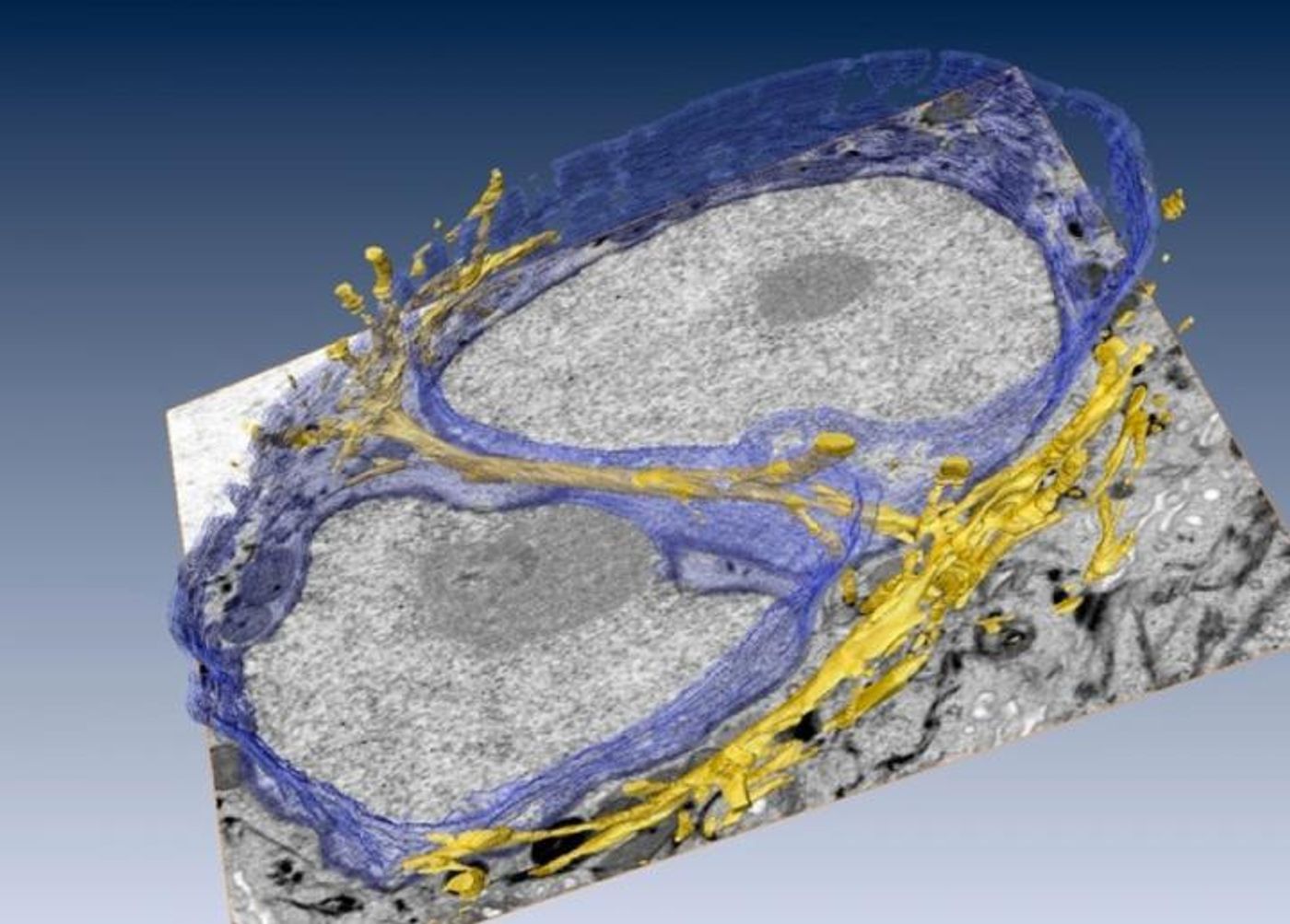
The blueprint for our bodies is contained in our genes, but scientists have long wondered how our environment can in turn exert an effect on our genes. It was first proposed in the early 80s by Mina Bissell of the Department of Energy's Lawrence Berkeley National Laboratory (Berkeley Lab) that cell fate and gene expression are influenced by a cell’s microenvironment or extracellular matrix. Bissell’s lab has now produced images that capture cytofilaments extending into the nucleus of a human breast cell. This research, published in the Journal of Cell Science, has demonstrated that it may be possible for the physical environment to impact the gene expression of a cell. Below is a three dimensional rendering of the cytofilament bundles tunneling through the nucelus of a cell, by Manfred Auer of the Berkeley Lab.
"There are somewhere between 30-70 trillion cells in our bodies, all with the same DNA sequence, so I've been saying since 1981 that something other than the sequence of the genes had to allow a nose to be a nose and not an elbow," said Bissell, co-corresponding author of the work and Distinguished Scientist at Berkeley Lab's Biological Systems and Engineering Division. "When the shape changes, biology changes."
Genes are generally seen as the primary drivers of cellular function, working with signaling molecules like hormones and transcription factors. The idea that the extracellular matrix (ECM) influences cellular processes has gained acceptance over many years of research. Work with breast cells of mice grown in a three-dimensional suspension supplemented with ECM showed that the right environment can influence cell behavior, in this case coaxing the cells to make milk.
"We knew the extracellular matrix was affecting gene expression, but it wasn't understood until now that the cytoskeleton was actually able to connect inside the nucleus," said Bissell. "Now we know there's a direct connection to the nucleus. That's what we're showing here for the first time. This is absolutely novel."
Bissell worked with Berkeley Lab’s Manfred Auer, head of the Cell and Tissue Imaging Department in the Molecular Biophysics and Integrative Bioimaging Division, who is co-corresponding author of the report.
"It took advances in cryogenic sample preparation techniques and large-volume electron microscopy to come up with these images," said Auer. Another special technique, stochastic optical reconstruction microscopy, or STORM, was applied to this research by another study author Ke Xu, a Berkeley Lab faculty scientist and UC Berkeley Assistant Professor of Chemistry.
"We combined a record-breaking six different imaging techniques and hundreds of millions of data points to obtain these images," said Auer. "The integrative bioimaging approach involved three different optical light and three different electron microscopy imaging approaches, each with its own strengths. This new integration of imaging approaches is what allowed us to study something as complex as this cytofilaments system."
The incredible resolution of their imaging allowed the researchers to show that the cytoskeleton associates with a class of protein which functions in the connection between the nucleus and cytoplasm of a cell, SUN proteins. The detail in the images enabled detection of grooves and tunnels in the nucleus. Cytofilaments stuck to the membrane of the nucleus moved through the invaginations, creating an architecture of cytofibers extending from the ECM to and through the nucleus.
"This study establishes for the first time the long-postulated mechanical link between the cell's nucleus to adhesion complexes that allow communication with the surrounding extracellular matrix and other cells," said Auer.
"The reason we're excited is that it explains a whole lot of literature of how force and tension could be playing a role together with biochemical signals to bring about huge changes in a cell," concluded Bissell.
Above, this video features a talk with Bissell about the ECM and its influence on the cell.
Sources: AAAS/Eurekalert! via Berkeley Lab, Journal of Cell Science