Barcoding Reveals Details of Blood Formation
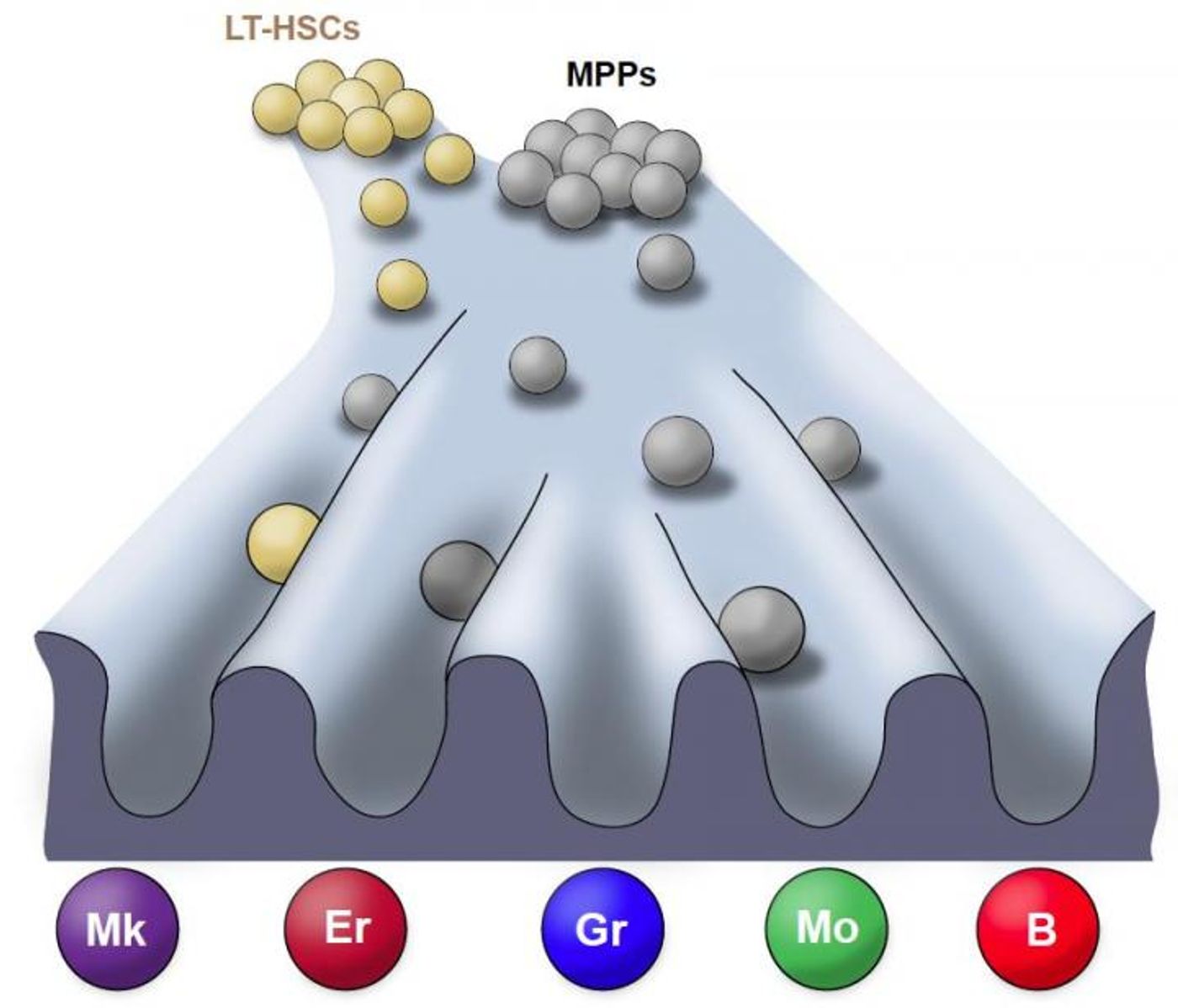
Following one cell through development can pose many challenges. Researchers overcame them by inserting tags into cells as a kind of barcode so they can be tracked individually in their natural state. In doing so, they observed that blood cells regenerate in a different way than how such cells behave following a transplant, calling into question some previous work. The data has been reported in Nature.
"The findings of this research, if applicable to humans, will have implications for blood cell transplantation, and for clinical and research methods using blood cells, such as gene therapy or gene editing," said John W. Thomas, Ph.D., Stem Cell and Cell-based Therapy Coordinator at NHLBI.
While this work is focused on the improvement of blood regeneration therapeutics, the researchers think it will apply to other cell types. It may yield insight about regenerative therapies that repair damaged or diseased tissues.
"Our results show that stem cells and their less pluripotent descendants, blood progenitors, behave somewhat differently when studied without removing them from their native environment versus when studied in a laboratory or in transplantation; leading to differences in the type of blood lineages they make," noted the first author of the work, Alejo Rodriguez Fraticelli, Ph.D., from Harvard Stem Cell Institute, at the Boston Children's Hospital.
Because there aren’t many useful techniques for the study of blood formation in the body’s natural state, most studies that have researched the lineage of individual blood cells have done so after a transplant. As such, the cells under investigation have been perturbed because they were excised from their normal environment. The scientists have suggested that their current models are a better representation of the directions that blood cells follow as they develop.
For investigator Rodriguez Fraticelli, blood regeneration is critical to study in its normal context. "Moving forward, we need to come up with methods to better predict what types of cells will be the most optimal for therapy, for instance in reprogramming cells, and editing," he said.
For this work, a transposon, which can insert in a random spot in the genome when an enzyme called transposase is present, was used to track the progenitors of blood cells as they move through the natural and unperturbed blood regeneration process. It has resulted in a revised roadmap that indicates how blood production and regeneration occurs in the natural state. The researchers also used single-cell RNA sequencing for their work.
An introduction to sequencing individual cells is presented in the video above from Illumina.
Sources: AAAS/Eurekalert! Via NIH/National Heart, Lung and Blood Institute, Nature