How Transcription Factors Find Their Target Sites
For an organism to function properly, genes have to be turned on or off in the right places at the right times. The machinery in our cells has a variety of ways to control gene expression, one of which are molecules called transcription factors. These proteins help regulate whether a gene gets transcribed into RNA, which must happen before it can be made into a protein or carry out some other function.
Transcription factors (which are described in the video) have to be able to first scan the genome so they can find their target sites and then bind there, which will turn genes on or off. It's known that they can also randomly attach to the genome non-specifically. When a transcriptions factor is more likely to randomly bind the genome, it becomes much better at finding its target sites because it can easily scan for it.
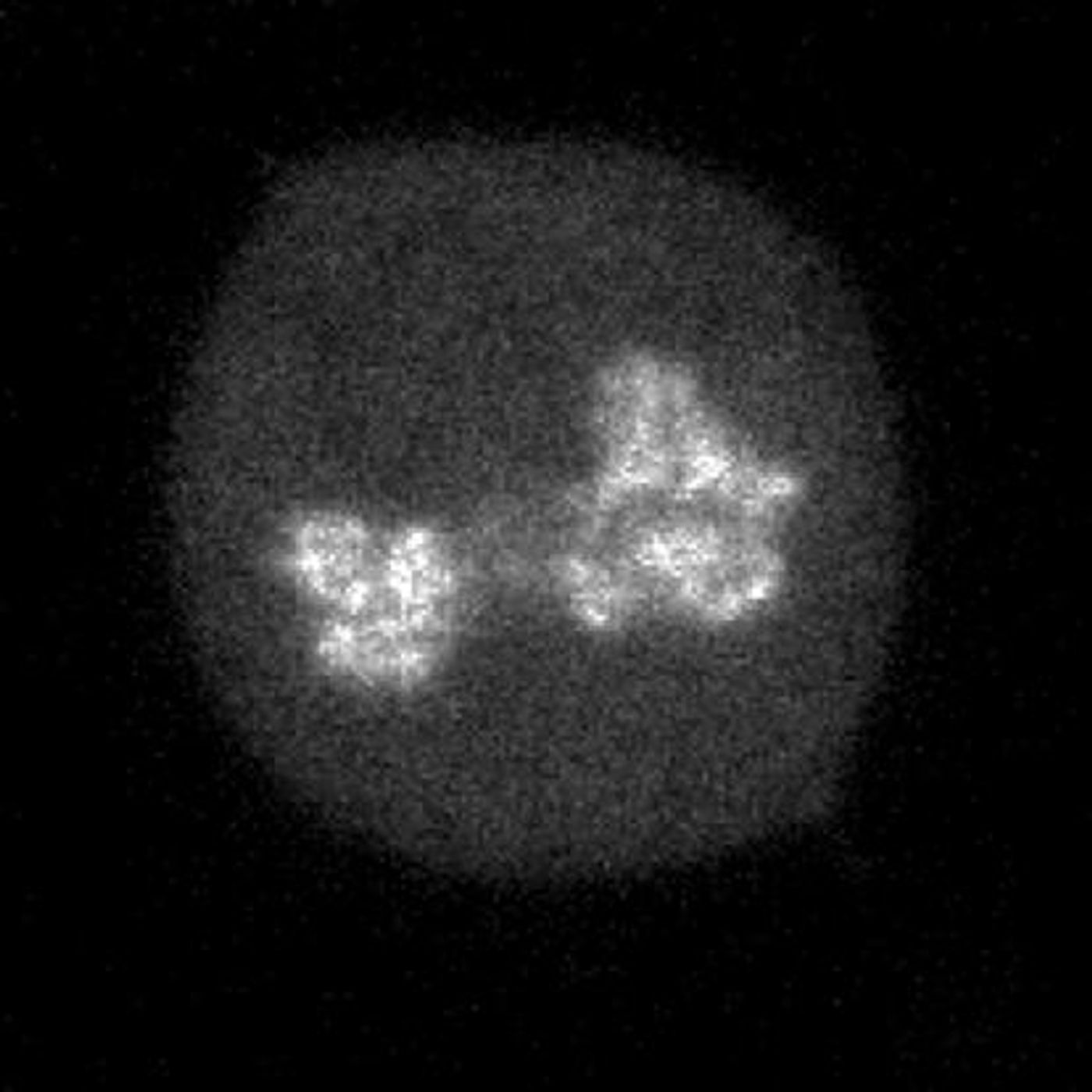
Researchers in the lab of David Suter at the Institute of Bioengineeering at the Ecole Polytechnique Fédérale de Lausanne (EPFL) have now developed a method for predicting how well different transcription factors look through the genome and find the place they need to be. They analyzed 501 mouse transcription factors by assessing how they attached to mitotic chromosomes. That characteristic has been connected to non-specific binding of transcription factors.
In this work, the team utilized photobleaching and single molecule imaging to demonstrate that the movement and efficiency of transcription factors can be estimated by mitotic chromosome binding.
"Transcription factors differ largely in their ability to scan the genome to find their specific binding sites, and these differences can be predicted by simply looking at how much they bind to mitotic chromosomes," explained Suter. "Transcription factors that are the most efficient in searching the genome could be able to drive broad changes in gene expression patterns even when expressed at low concentrations, and can therefore be particularly important for cell fate decision processes."
The researchers mapped the transcription factors in the genome and found that the ability of the molecules to find their target sites differed by three orders of magnitude. The work demonstrates that transcription factors that tend to bind to DNA in a non-specific way also move slowly in the nucleus, associate with mitotic chromosomes, and are good at finding their target sites.
Sources: AAAS/Eurekalert! Via EPFL, Nature Communications