Denisovan DNA Influences the Immune System of Oceanian People
Our DNA helps tell the story of human evolution. As species in the genus homo evolved, our ancient ancestors interbred with populations of Neanderthal (Homo neanderthalensis) and Denisovans (discovered in 2008, they don't have a scientific name yet). Many modern humans (Homo sapiens) still carry DNA from Neanderthals and Denisovans. While people have genomes that contain more or less the same set of genes, there are differences in the sequences of those genes that make us unique.
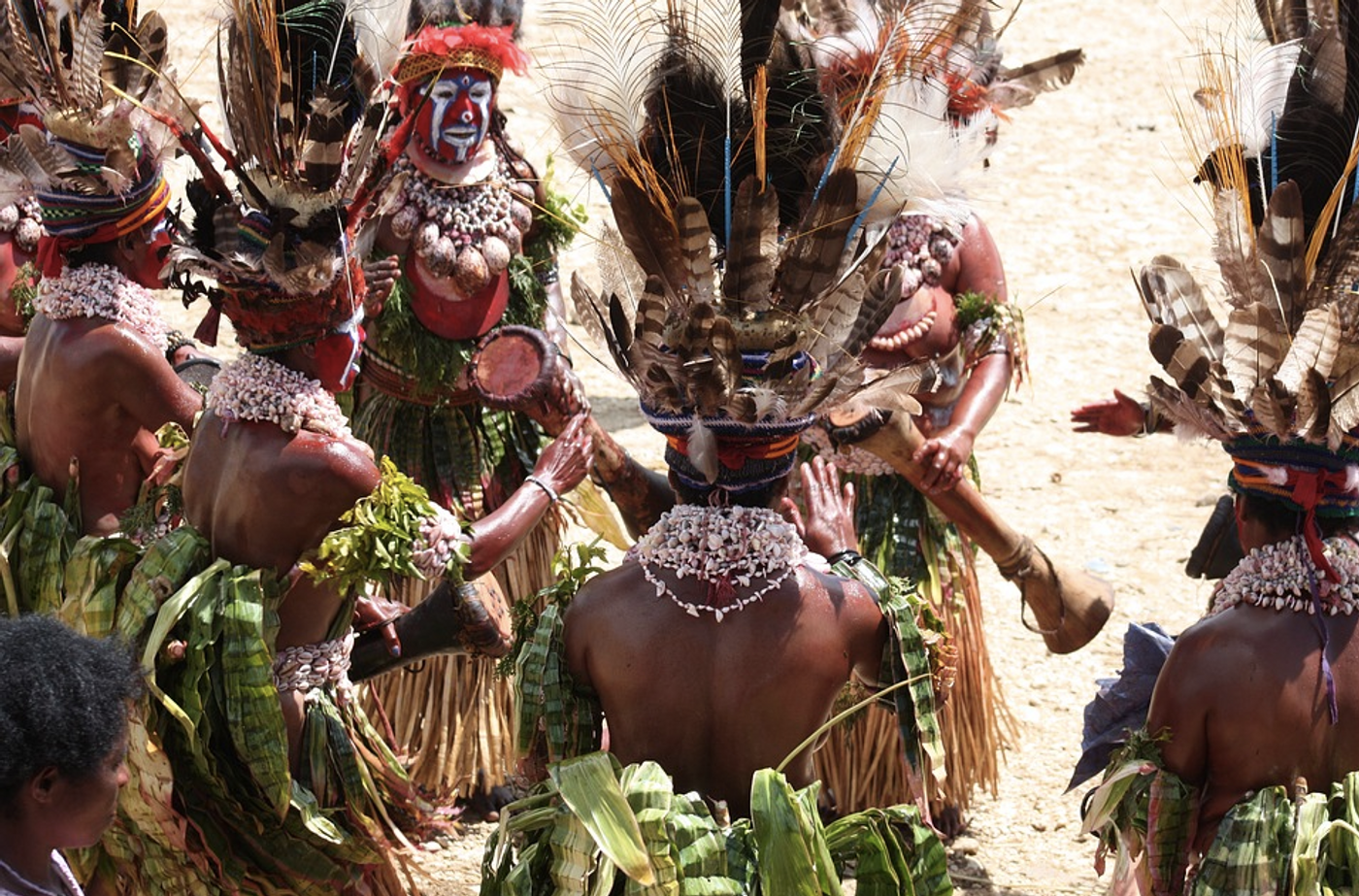
Researchers have now identified over 120,000 variations in the human genome that change the genomic structure, and in turn, they influence large genomic regions. Some of these regions are connected to disease susceptibility or the immune response. The study, reported in Cell, also showed that some people from Papua New Guinea carry genetic changes they inherited from Denisovans, which have caused physiologically significant alterations. These variations could have dramatic impacts on how well medical treatments work for some populations.
Many studies that have investigated how genetic changes influence biology or disease risk have focused on changes on individual bases of the genetic sequence. This study has broadened the approach, to look at changes involving a handful of bases to a few million.
"By analyzing the genomes of understudied populations we've been able to find high-frequency structural variations not uncovered by previous large-scale sequencing projects," noted the first study author, Wellcome Sanger Institute graduate candidate Mohamed Almarri. "Several of these are in medically-important genes that tell us how a population has evolved to resist a certain disease or why they might be susceptible to others. This is vital knowledge and will help to ensure that treatments can be tailored to each specific population."
The Wellcome Sanger Insitute has previously sequenced genomes from 911 people that were from culturally and geographically diverse groups from around the world. This study assessed the structural changes inherent in these sequences. Many that were identified were not previously known. One that was found is a deletion from a gene called AQR, which is involved in the detection of and immune response to viruses.
The scientists also found duplication events in which people have ended up carrying multiple copies of a gene. In all of the African populations that were studied, multiple copies of a gene called HPR were found; the gene is linked to resistance to sleeping sickness. In places where the disease is most common, like Central and West Africa, people carried the highest numbers of copies - up to nine were found.
"This is a very valuable study showing the importance of structural variation of the human genome in the genetic diversity of humans around the world. The work supports the concept that some human adaptations to different environments are due to the loss or gain of whole genes, or parts of genes," said Dr. Ed Hollox of the University of Leicester. "Structural variation can be challenging to find, and this study also provides a well-founded structural variation reference set which will serve as an important springboard for future studies."
This research is an important addition to the human reference genome, which has been limited in what it can tell scientists because it was created from only a handful of people. If those people did not carry some variation, it's not in the reference genome. A comparison of these newly sequenced genomes and the reference genome shows that we still have a lot to learn about the diversity in the human genome.
"Structural variants are complicated yet very important functionally, evolutionarily, and medically. The discovery of these new structural variations provides one of the richest resources of this kind of variation so far, which not only offers unique insights into population histories and improves the currently used human reference genome, but will also substantially benefit future medical studies," added Dr. Yali Xue, who has recently retired from the Wellcome Sanger Institute.
Sources: Phys.org via Wellcome Trust Sanger Institute, Cell