Deciphering a Massive Genome
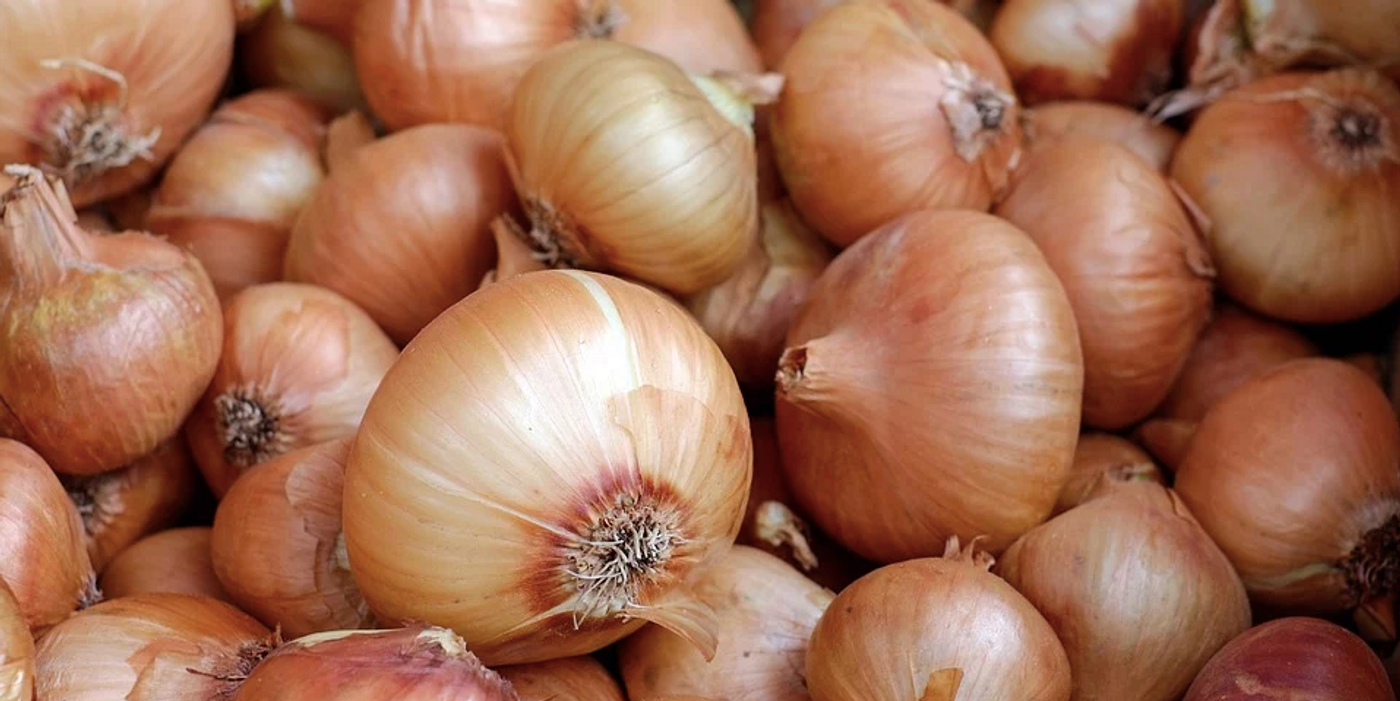
The human genome is about 6 billion base pairs long, the Escherichia coli bacterium's genome is about 5 million base pairs long, and the onion has a genome that is around 16 billion base pairs in length. The onion genome is not nearly as long as the largest known, that designation belongs to the Japanese flower Paris japonica; it's about 149 billion base pairs. But the length and characteristics of the onion genome have made it very difficult to sequence.
“In this study, we challenged the limits of how far we can go in deciphering a DNA sequence consisting of 16 billion base pairs,” said study author Masayoshi Shigyo, a professor in the Graduate School of Sciences and Technology for Innovation at Yamaguchi University.
When a genome is mapped, scientists can usually work with a reference that has been assembled from the genes of that species. Sections of the genome from different organisms can be compared to others of the same species or those that are closely related. But the onion genome is so large that it had to be combined with a different approach.
“In previous work, we collected the sequence information on expressed genes, determined the locations of 25,000 expressed genes and succeeded in developing the world’s first high-density genetic map that accurately reflects the order of expressed genes,” Shigyo said.
Then, the researchers extracted DNA from onions grown in the lab to create more genetic sequence data. These sequences were compared to a genetic map that was used to represent onion chromosomes. This represented about half the genetic material in an onion plant. Chromosomal sequences were used to make predictions about the number of genes in the onion genome,
“The prediction method detected 540,925 putative possible genes,” Shigyo said. “This number was much higher than expected, suggesting there are many pseudogenes in the genome.” Pseudogenes are a kind of non-coding or 'junk' DNA, Shigyo said.
Genes with a clear function were distributed evenly across the genome.
“In short, due to the large amount of junk DNA in the genome, computer sequence assembly cannot be accomplished,” Shigyo said. “However, in these difficult circumstances, our high-density genetic map information served as a model, and we succeeded in beginning to decode the onion genome for the first time.”
This work could help onion growers create better cultivation methods, said Shigyo.
“Humans have a long history of cultivating onions, dating back to 2300 BC in Egypt, and the vegetable is believed to have contributed to our health and society since then,” Shigyo said. “Yet onions tend to be thought of as suitable for high school biology experiments, but not more serious research. I hope the results of this work will bring more attention to onions, and the potential benefits of better understanding the crop, in the research community.”