First, there was the Human Genome Project, a monumental, international effort to map the entire human genome down to its base pairs of As, Ts, Cs, and Gs. Following this, other projects have arisen to explore all of the biological complexities of the human race. Fundamental biologists have focused on the central dogma of molecular biology, the process of turning DNA into RNA into functional products, like proteins. A new project proposed this week sets out to explore a new group: human proteoforms.
A proteoform is a simple term that describes protein complexity, explains Lloyd M. Smith, the first author of the new review published in Science Advances. More explicitly, proteoforms are all versions of a single protein— its RNA splice variants, posttranslational modifications, even its potential genetic polymorphisms. All forms of a protein make up its proteoform (hence the name).
The paper is titled The Human Proteoform Project: Defining the human proteome, and it lays out the necessary steps and goals of this new undertaking. The project’s primary goal is to map out and define the proteoforms defined from the approximately 20,000 genes in the human genome. This is a two-pronged approach: first, the plan is to investigate medically relevant proteoforms, ones important to diseases; second, scientists will invest and research other technologies that will allow for large-scale analysis of healthy proteins.
Researchers drew inspiration from the Human Genome Project, which began with a framework for the human genome while continuing to advance technology to sequence the entire genome efficiently. This project is also done in synergy with other initiatives, like the Human Proteome Project and the Human Protein Atlas, two well-established endeavors to define all of the proteins in the human body. While guiding the Human Proteome Project, the Human Proteome Organization has called for community-led initiatives to map the human proteoforms as well.
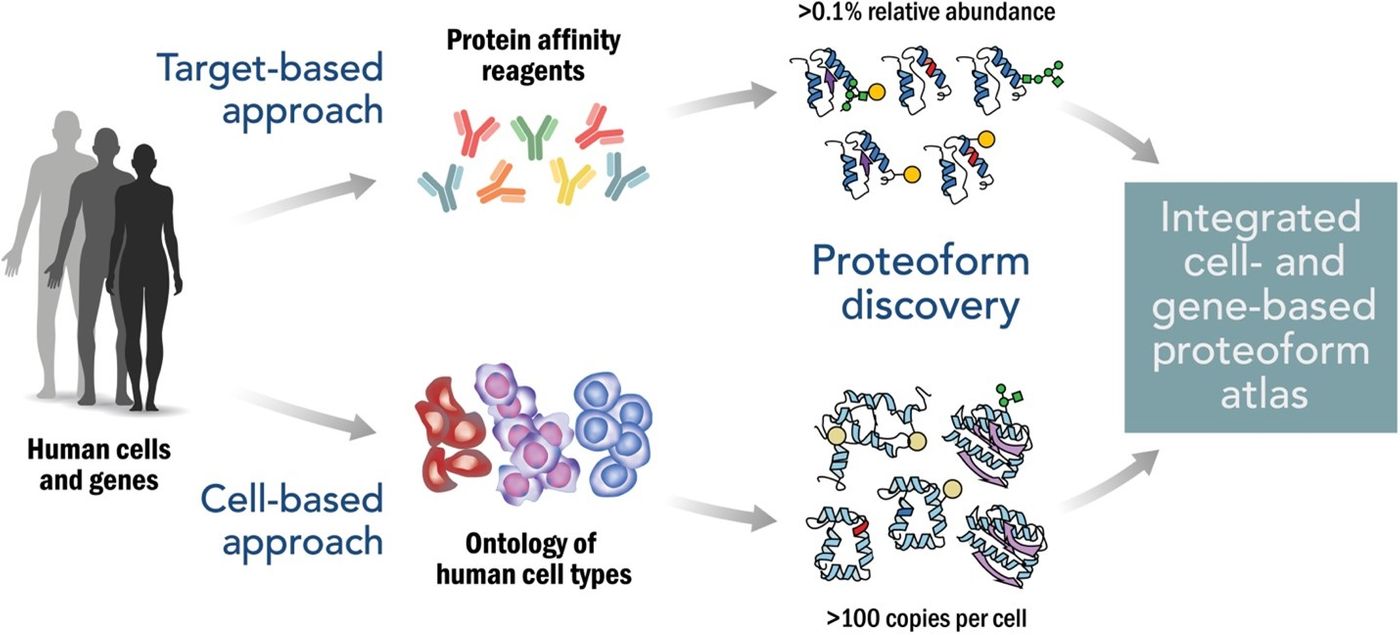
The result of the Human Proteoform project will be The Human Proteoform Atlas, assembled using both a target-based approach and a cell-based approach. The target-based method uses protein affinity reagents such as antibodies to isolate proteoform families from specific genes. The cell-based process will be more expansive, focusing on cell types to characterize specific proteoforms. Researchers note that emphasis will be given to identifying the “dominant proteoform population” instead of rare occurrences.
Ultimately, the authors of this paper believe that this project will enable critical breakthroughs in biomedical research, including regenerative biology, drug development, and the detection of human disease. They also hope that their investment in technology for this project will help advance other fields of biology and chemistry. There is a lot to look forward to with this exciting new venture, and we will all be watching closely for updates!
Sources: The Human Genome Project, Science Advances