Needle in the haystack: How to remove human background when you want to detect microorganisms
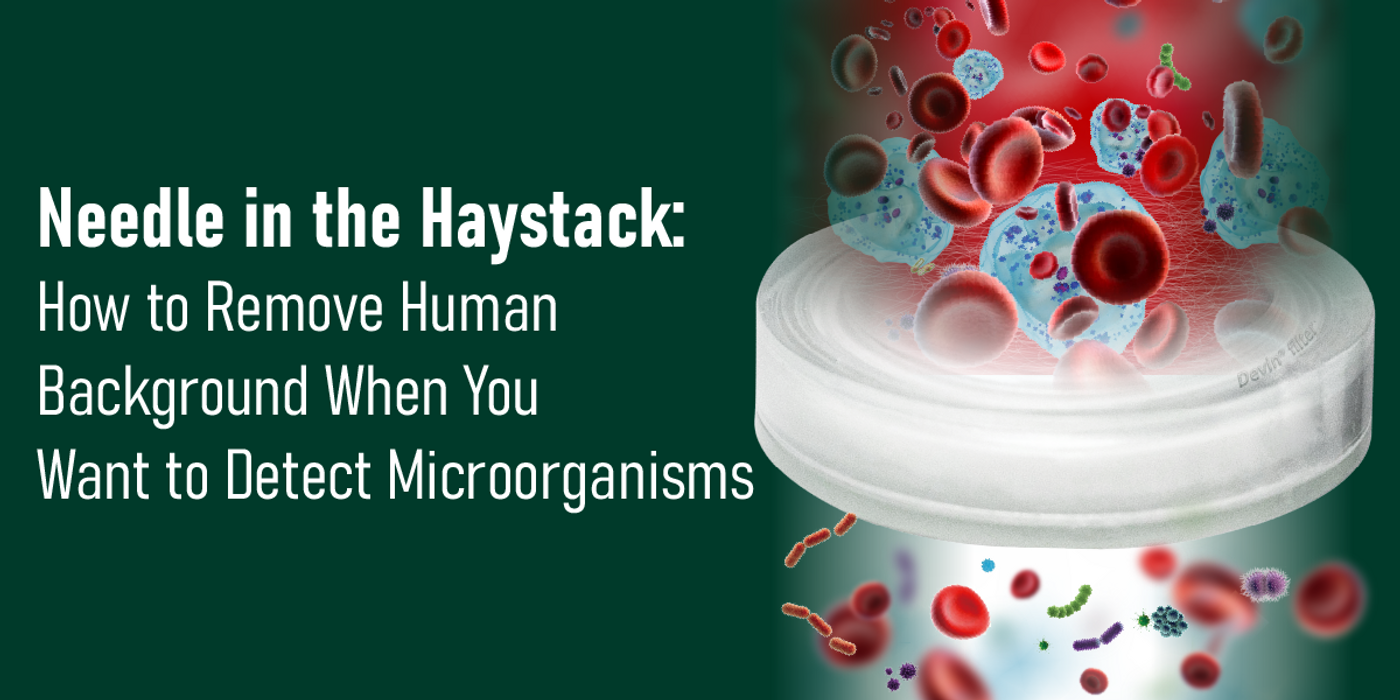
To substantially reduce the risk of death, accurate and timely identification of the causative pathogens is crucial for selecting the proper treatment for the patients.
Currently, blood culture is still the gold standard test to identify bloodstream pathogens. However, this process can take 2-7 days, with the positivity rate less than 20%, which means in over 80% of the cases, the pathogens remain unidentified.
However, to detect the extremely low amount of microorganism, host DNA interference is still one of the major obstacles to use sequencing technology in clinical microbiology.
Host DNA interference >99%1 is major bottleneck in clinical microbiology.
In this article we analyzed most recent techniques available on the market, compared their efficacy and outlined perspective of human DNA depletion for mNGS application in clinical settings for rapid pathogen detection and timely treatment of infectious diseases.
From this review you will learn which depletion methods prior or post DNA extraction are available, will get to know advantages and drawbacks of each method and will be introduced to a novel technique PaRTI-Seq®, developed by Micronbrane Medical that deplete >95% of human leukocytes from whole blood in less than 5 minutes and enables identification of pathogens within 24 hours.
1 A key finding of the whole metagenome shotgun sequencing of different sample types conducted by the Human Microbiome Project