Scientists have discovered that microbiota that colonize our bodies inside and out have the capacity to create molecules that function as drugs-molecules that may one day help pave the way for new, synthetically manufactured drugs to cure what ails us.
These drugs, which occur spontaneously in the human body, may play a role in keeping us healthy. "We used to think that drug s were developed by drug companies, approved by the FDA, and prescribed by physicians, but we now think there are many drugs of equal potency and specificity being produced by the human microbiota," says Michael Fischbach, PhD, assistant professor of bioengineering, University of California San Francisco School of Pharmacy, who routinely discovers compelling molecules made by microbes.
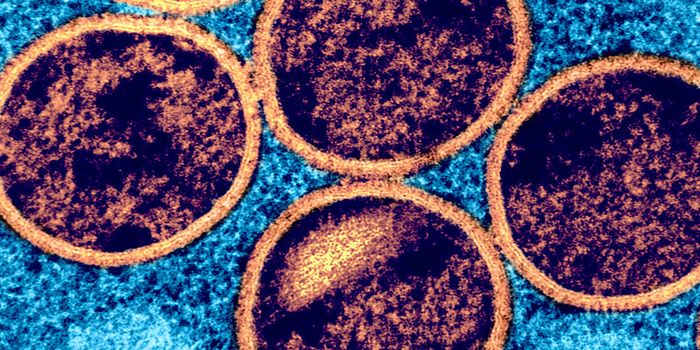
A recent example can be found in a study led by Fischbach, published in the journal Cell. Investigators purified and deciphered the structure of one of the molecules they identified-an antibiotic they named lactocillin, which is made by a common bacterial species, Lactobacillus gasseri, found in the microbial environment within the vagina. Lactocillin discriminately obliterates certain vaginal bacterial pathogens while bypassing harmless species. Pharma companies are clinically testing antibiotics that resemble lactocillin. The scientists showed that lactocillin has strong antibacterial activity against a variety of Gram-positive vaginal pathogens.
Fischbach notes that about one-third of all therapeutic drugs-for treating bacterial infections, cancer, and high cholesterol-have been derived from microbes and plants. While microbes found in the far reaches of the earth have been a potent source material for drug makers, a prized new source may be closer to home. People harbor hundreds of distinct species in different parts of their bodies.
In addition to describing the microbiomes in our gut, on our skin, inside our nose, mouth, and vagina, scientists are finding microbiomes where the range and amount of species diversity diverges from what's normal to aspects linked with disease risk factors.
Fischbach's team has made use of new data-analysis software that it created on a comprehensive genetic database, identifying clusters of bacterial genes that are switched-on in an organized way to guide the production of molecules that are biologically active in people. ClusterFinder, the algorithm the team designed, draws conclusions from new data that builds on known links between gene clusters in soil and ocean-going bacterial species and molecules they produce.
With ClusterFinder, the team was able to analyze genomes from microbiome species and data on gene activity from human samples to identify more than 3,100 distinct clusters of bacterial genes found throughout the body. These clusters encode enzymes that synthesize drug-like molecules that fit into pre-existing classes of medicines.
Fischbach says the study shows that genus-level analysis that is routinely relied on to identify bacteria in human microbiomes is not sufficiently intricate to foresee which drug-like molecules the bacteria produce, as particular species and varies strains within each of them create different molecules. "We need to learn what these molecules are and what they are doing," he says. "This could represent a pool of molecules with many tantalizing candidates for drug therapy.
The scientists are drilling down, literally. "It's been clear for several years that variations and changes in the human microbiome have interesting effects on the human host, and now we can begin to determine why this is true on a molecular level," Fischbach says.
 Scientists have discovered that microbiota that colonize our bodies inside and out have the capacity to create molecules that function as drugs-molecules that may one day help pave the way for new, synthetically manufactured drugs to cure what ails us.
Scientists have discovered that microbiota that colonize our bodies inside and out have the capacity to create molecules that function as drugs-molecules that may one day help pave the way for new, synthetically manufactured drugs to cure what ails us.