A
new study published in Science has shown that the bacteria in our guts has been with us a long time, and coevolved alongside hominids. This shows that the community of bacteria in our gastrointestinal tract, the microbiome, is not just picked up from our environment and altered by medicine or certain foods but is also influenced by our evolutionary history. It also suggests that the microbiome has aided in shaping our digestive tract. If you want to know more about the gut microbiome itself, check out the video below from Mayo Clinic, and find out about some of the latest research connecting the microbiome to human disease.
"We know that gut bacteria are really integrated with our biology, and this research gives us a framework for investigating how that has come to be," said Andrew Moeller, a Postdoctoral Fellow at the University of California, Berkeley's Museum of Vertebrate Zoology. Moeller led the work along with Howard Ochman, a Professor of Integrative Biology at The University of Texas at Austin.
"It's surprising that our gut microbes, which we could get from many sources in the environment, have actually been co-evolving inside us for such a long time," said Ochman, also noting that the bacterial flora have been propagated over hundreds of thousands of generations of hosts.
To carry out this study, Moeller and colleagues used a new approach. Typically, analysis of the microbiome takes advantage of a particular piece of a bacteria’s genome, the 16S ribosomal subunit. That specific part of bacterial DNA doesn’t change much and as such easily indicates the identity of the bacteria in the community when it’s sequenced. Because of that lack of change, the team switched to a gene that has a higher rate of evolution, called gyrase B (it encodes for DNA gyrase - a protein that helps DNA uncoil and coil).
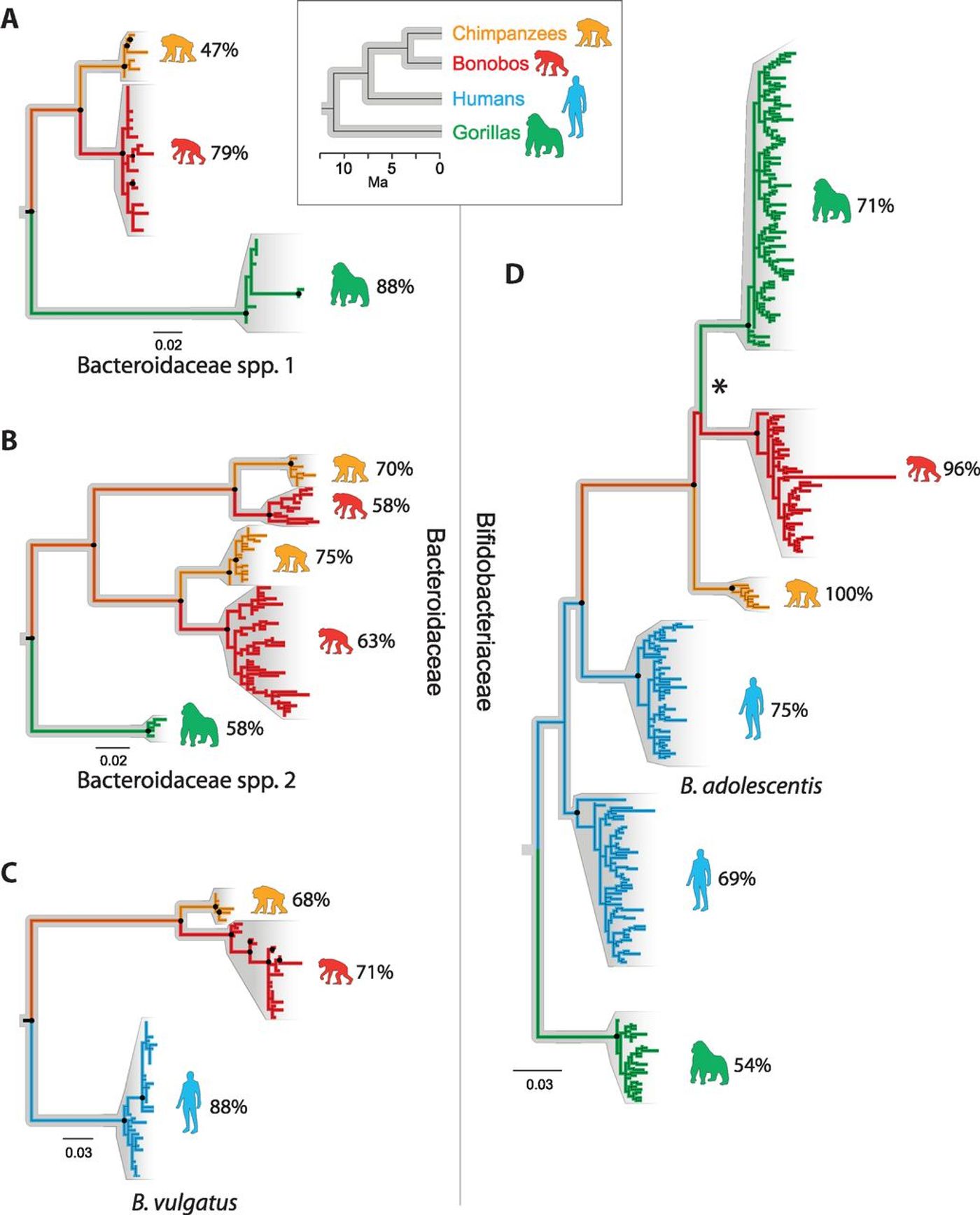
Using fecal samples obtained from the environment of the hosts under study, bacterial DNA was isolated from chimps in Tanzania, gorillas in Caneroon, bonobos in the Democratic Republic of the Congo, and humans in Connecticut. A microbiome family tree was constructed for three common and abundant human gut bacteria - Bacteroidaceae, Bifidobacteriaceae and Lachnospiraceae.
The investigators found that only the Bacteroidaceae and Bifidobacteriaceae families showed cospeciation with their ape hosts; gut microbial diversity was highest in gorillas but lowest in humans.
"Once we calibrated the molecular clock, we were able to date the split of human and chimp bacteria at around 5.3 million years ago, and the human-gorilla gut bacteria split at around 15.6 million years ago, which are roughly in line with what we know from fossil and genomic data of the hosts," Moeller explained. "It is one more line of evidence that gut bacteria have cospeciated with humans."
"In a way, host speciation is like continental drift: When two continents drift apart, whole biotas begin to diverge," Moeller continued. "Here, as the hosts are splitting, a good chunk of their microbiota is also splitting and diversifying."
Other findings show that the Lachnospiraceae tree was more complicated. There were propbably at least four times when bacteria in this family got transferred to a different host species, something the team suggests was easy for these bacteria since they form spores.
"We have shown that the microbiome is a composite of microbial lineages, some that have cospeciated with us, and some that have been passed around from one host species to another," explained Moeller.
A comparison of the microbial flora of Africans from Malawi to the U.S. human microbiota showed distinct differences, with some lineages apparently lost by Americans.
"It will be interesting in the future to do a full study of human populations using this strain-level method to see whether we can use bacteria to reconstruct the history of human migrations,” said Moeller. “What's most exciting to me is the possibility that this codiversification between bacteria and hosts could extend much further back in time. Maybe we can trace our gut microbes back to our common ancestors with all mammals, all reptiles, all amphibians, maybe even all vertebrates. If that's true, it's amazing," he concluded.
Sources:
Science Daily via
University of California, Berkeley,
Science