Rising antibiotic resistance is considered a
major public health threat; one that the
United Nations has taken notice of, prompting warnings about the danger it poses. Utilizing computational power, researchers looked to bacteria to find candidate compounds they were then able to synthesize. Two potential new antibiotics have been produced using this strategy.
A research team at Rockefeller University led by Sean Brady, head of the
Laboratory of Genetically Encoded Small Molecules, interrogated public databases containing the genomes of bacteria that live in the human body. They wanted to find clusters of genes that were predicted to produce molecules, non-ribosomal peptides, which are the basis of myriad antibiotics.
There are genes within the genome of a microbe that are antimicrobial, but using bacteria to produce those compounds is fraught with challenges. Many bacterial strains are difficult to grow in a lab. Scientists would also have to determine how to manipulate the proper genes of a bacterium to make them express the antimicrobial molecules. Using computational methods avoided those problems.
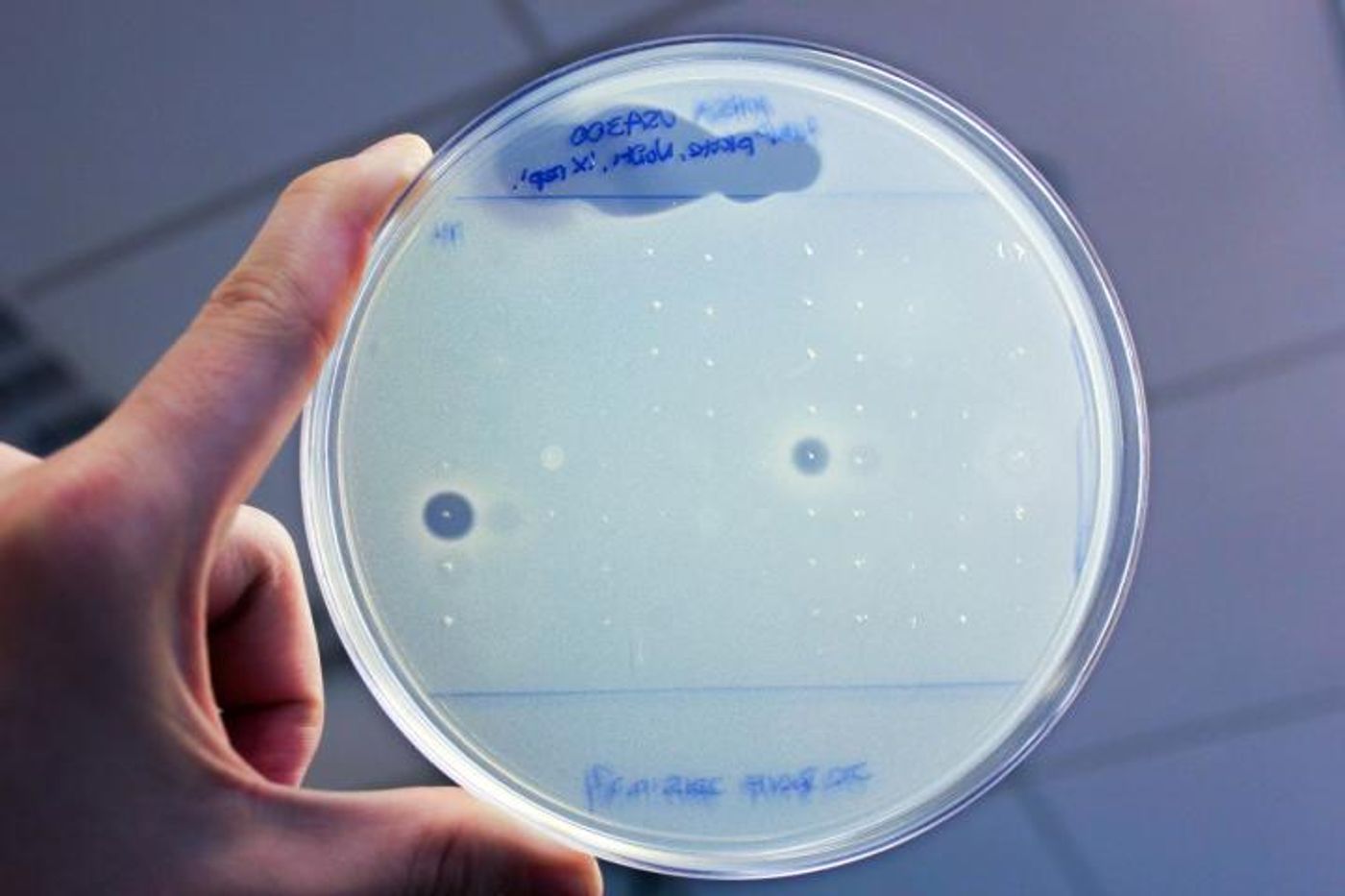
In doing so, the team found 57 candidate gene clusters, from which 30 were chosen. Using a technique called solid-phase synthesis, 25 compounds were produced. Those compounds were then tested with human pathogens, enabling the investigators to identify the two antibiotics the scientists named humimycin A and humimycin B.
The findings were published in Nature Chemical Biology.
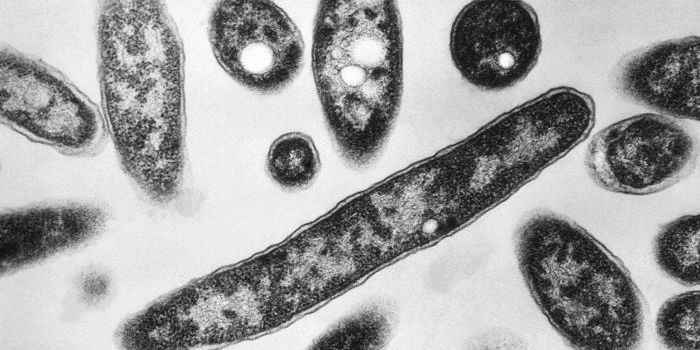
Humimycin A and B are closely related and from a family of bacteria named Rhodococcus, which have not previously yielded anything resembling the humimycins when grown with typical laboratory protocols. The humimyics proved very effective combating Staphylococcus and Streptococcus bacteria, microbes that can result in serious infections in humans and tend to become antibiotic resistant.
Experiments determined that the humimycins act to inhibit an enzyme that aids in buildings cell walls in bacteria; without that cell-wall-building power, the bacteria die. Beta-lactams are a commonly used class of antibiotics that have a diminishing effect as bacteria build resistance to them. The researchers found, however, that humimycins can actually re-sensitize bacteria to the action of beta-lactams they had become resistant to.
Staphylococcus microbes that were resistant to beta-lactam died after being exposed to a beta-lactam antibiotic in combination with humimycin A. Incredibly, that effects was still seen even if there had been little effect from using humimycin A by itself. Brady suggested that is because both compounds work by disrupting different parts of the same biological pathway.
"It's like taking a hose and pinching it in two spots," he says. Even if neither kink halts the flow altogether on its own, "eventually, no more water comes through."
The scientists went a step further and used a common cause of antibiotic-resistant infections, Staphylococcus aureus, to infect mice. After being dosed with the strain, which was beta-lactam resistant, the mice were treated with a either humimycin A, a beta-lactam antibiotic, or a combination of both. The mice treated with both drugs had the best outcome, and that discovery could pave the way toward therapeutics for human use.
Brady is hopeful that this work will inspire others to pursue a similar line of reasoning to find other potential therapies. He plans to apply the strategy to bacterial species outside of the human microbiome, and eventually, even to those bacterial species whose genomes have not yet been sequenced.
Sources:
Pharmacy & Therapeutics,
United Nations,
AAAS/Eurekalert! via
The Rockefeller University,
Nature Chemical Biology