Genomic Sequencing Gives Insight Into Shigella Outbreaks
Researchers at UC Davis and the California Department of Public Health (CDPH) have spearheaded an effort to apply genetic sequencing tools to help improve public health. The investigators used samples of Shigella sonnei bacteria that was associated with major shigellosis outbreaks in California in recent years, sequencing the bacterial genomes. Learn more about shigellosis in the following video.
S. sonnei is one of four types of Shigella and causes most outbreaks of shigellosis, resulting in gastrointestinal problems like abdominal pain and diarrhea. Shigellosis causes around 500,000 infections, 6,000 hospitalizations and 70 deaths in the United States every year.
The data obtained has revealed the presence of antibiotic resistance genes, gives some insights into the pathogenicity of the bacteria, and illustrates how the California strains are related to others around the world. The scientists have published the first major, whole-genome analysis of North American S. sonnei strains in mSphere.
"If you have an outbreak and you want to know what is causing a particular problem, like antibiotic resistance, sequencing the genome can identify the genes involved," said study collaborator Jonathan Eisen, a professor with appointments in the Department of Medical Microbiology and Immunology and the Department of Evolution and Ecology at UC Davis. "Eventually, we should be able to sequence whole genomes of bacteria to support patient care," he said.
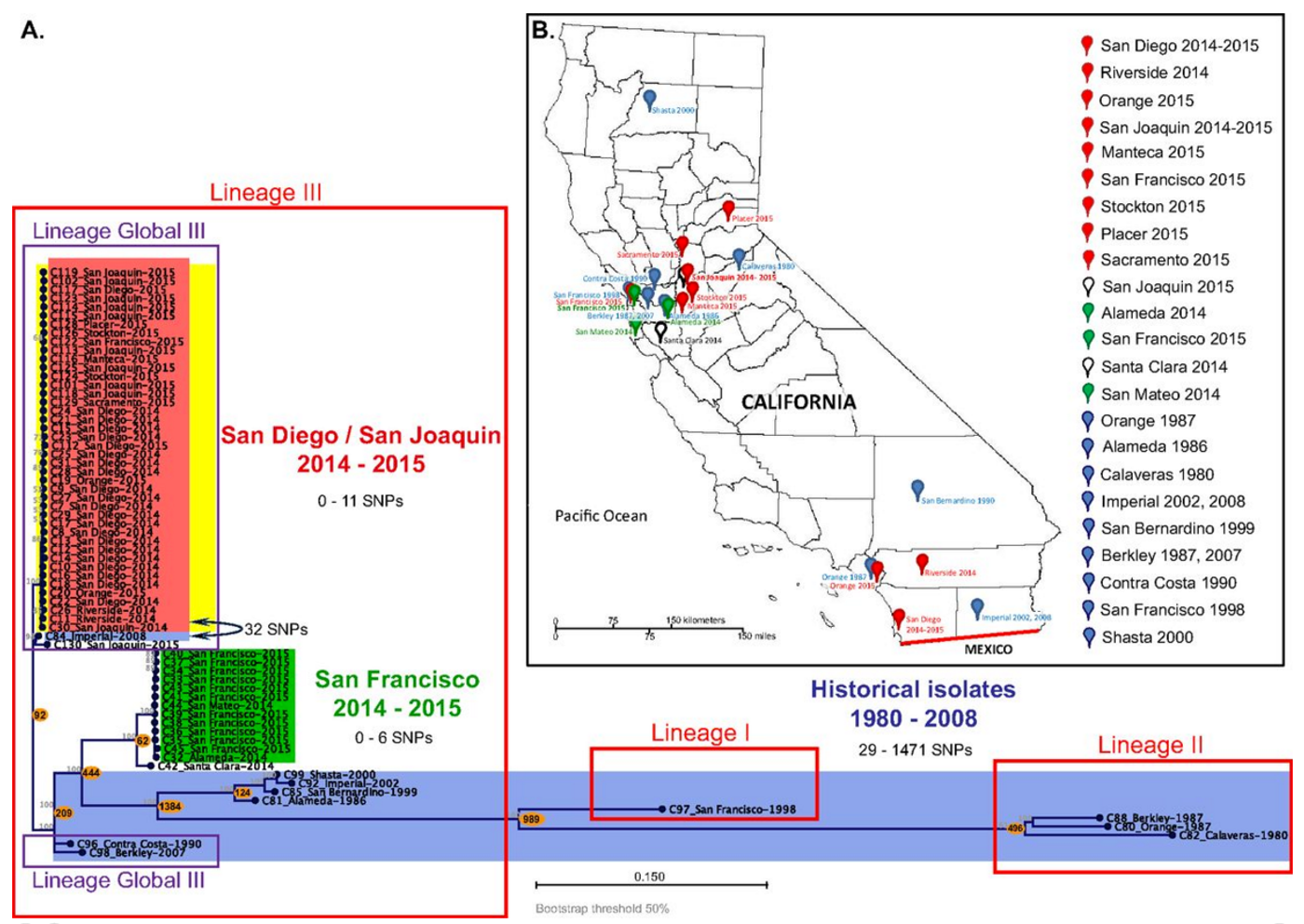
For this study, samples from the recent California outbreaks and historical strains from California, Asia were included in a group of 68 isolates whose genomes were sequenced and assayed for antibiotic resistance.
The researchers found two clusters in these outbreaks, one strain that primarily struck San Diego and the San Joaquin Valley, which has been in California for at least nine years. Another strain caused and an outbreak localized to the San Francisco Bay Area. However, it was discovered that some of the isolates were infected with a virus that attacks bacteria, a bacteriophage. In that bacteriophage was a Shiga toxin (STX) gene, identified in the more dangerous S. flexneri and S. dysenteriae. Check out a news report on the Bay Area outbreak below.
"Shigella sonnei bacteria normally cause a less severe disease and are not known to produce Shiga toxin," explained Dr. James Watt, Chief, Division of Communicable Disease Control at CDPH. “The toxin gene was most likely acquired by Shigella sonnei via genetic exchanges with E. coli and other Shigella species. Discovering a functional toxin gene was concerning in this large outbreak. Finding this gene raises concerns that illness due to Shigella sonnei could become more severe in the future," Watt said.
The strain affecting San Francisco did not carry STX but instead, had genes that conferred resistance to fluoroquinolone antibiotics. Because the fluoroquinolone-resistance genes had similarities to strains from Southeast Asia, it is likely a vital clue as to the origin of the strain.
"We know these movements of DNA can be important for the spread of antibiotic resistance, virulence and pathogenicity factors," Eisen said. "Having the genome data from outbreaks allows us to try to figure out what happened."
The researchers suggest that studies like this one would benefit patients; “knowing the variants of a bacterial strain would only improve therapeutics. “If you can show that the transfer of antibiotic resistance genes came in response to some kind of treatment, you would certainly think about isolating people who were receiving that treatment, potentially sampling them more often or even changing treatments," explained Eisen.
It is time to apply the powerful genomic tools we have at hand to public health. “It is clear, from a technical, economic, and scientific point of view, that we can and should be integrating more genome sequencing into infectious disease studies," Eisen concluded.
Sources: AAAS/Eurekalert! via UC Davis, mSphere