Gut microbes talk to the immune system
Researchers from Harvard Medical School have reported on the results of a large-scale search for immunomodulatory organisms. Specifically, they analyzed how gut bacteria affect immune cells and gene expression.
These findings will help researchers design individualized treatments for different diseases based on how the gut microbiome interacts with the immune system. According to study author Dennis Kasper, “we set out to map out interactions between bacteria and the immune system in the hope that this could eventually lead to the development of an apothecary of agents tailored to modulate the immune system selectively and precisely.”
The group collected 53 of the most common human gut bacteria and transferred them individually to germ-free mice; these 53 species represented the five dominant phyla of gut bacteria: Bacteroidetes, Firmicutes, Proteobacteria, Actinobacteria, and Fusobacteria. Once the mouse guts were colonized with one of the species, they analyzed how each bacterial species affected immune cells and gene expression.
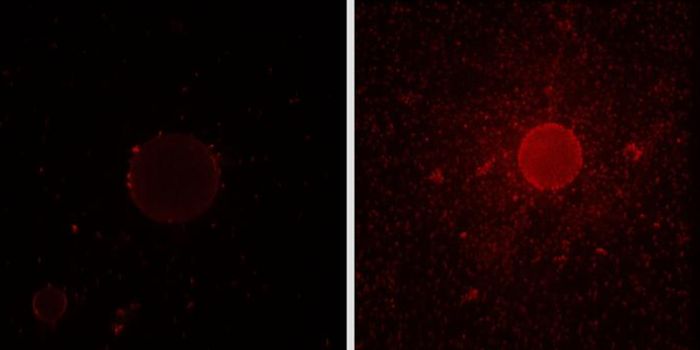
Over one-third of the bacterial species they tested increased the number of plasmacytoid dendritic cells in the colon - by at least two-fold. Plasmacytoid dendritic cells are innate immune cells that help detect pathogens. In contrast, some species - Staphylococcus saprophyticus and Lactobacillus rhamnosus - decreased the number of these immune cells (actually, these mice had virtually no plasmacytoid dendritic cells).
With respect to T cells, Fusobacterium varium decreased the number of CD4+ and CD8+ αβT cells.
Not only did various species of bacteria alter the populations of certain immune cells, they also altered gene expression. They found that Bacteroides dorei and B. longum both induced innate lymphoid cells in the gut to produce IL-22, a cytokine that helps initiate the innate immune response against bacterial pathogens. However, some bacteria - Acinetobacter lwoffii, Clostridium sordellii, and Veillonella - prevented the production of IL-22.
Lastly, certain bacteria altered the production of antimicrobial peptides in the gut. F. varium downregulated the expression of the Reg3 genes. On the other hand, Parabacteroides merdae and Porphyromonas uenonis induced the production of α-defensin, but decreased the expression of Reg3.
According to Kasper, “we believe that some microbes may upregulate certain genes to create a more hospitable environment for themselves, while others may downregulate certain ones to create a more hostile one for harmful bacteria.”
Sources: Cell, Science Daily, Wikipedia