A new Tool to Identify Bacterial Infections
It can be incredibly challenging to give a patient the proper treatment when they are infected by an antibiotic-resistant pathogen. For those living in poor or rural regions, it can be even more difficult because of a lack of reliable infrastructure, and limited access to quality diagnostic labs that have specialists with the expertise to figure out what exactly is ailing people. Currently, doctors have to isolate the pathogen and grow it with an assay. That process can take hours or even weeks.
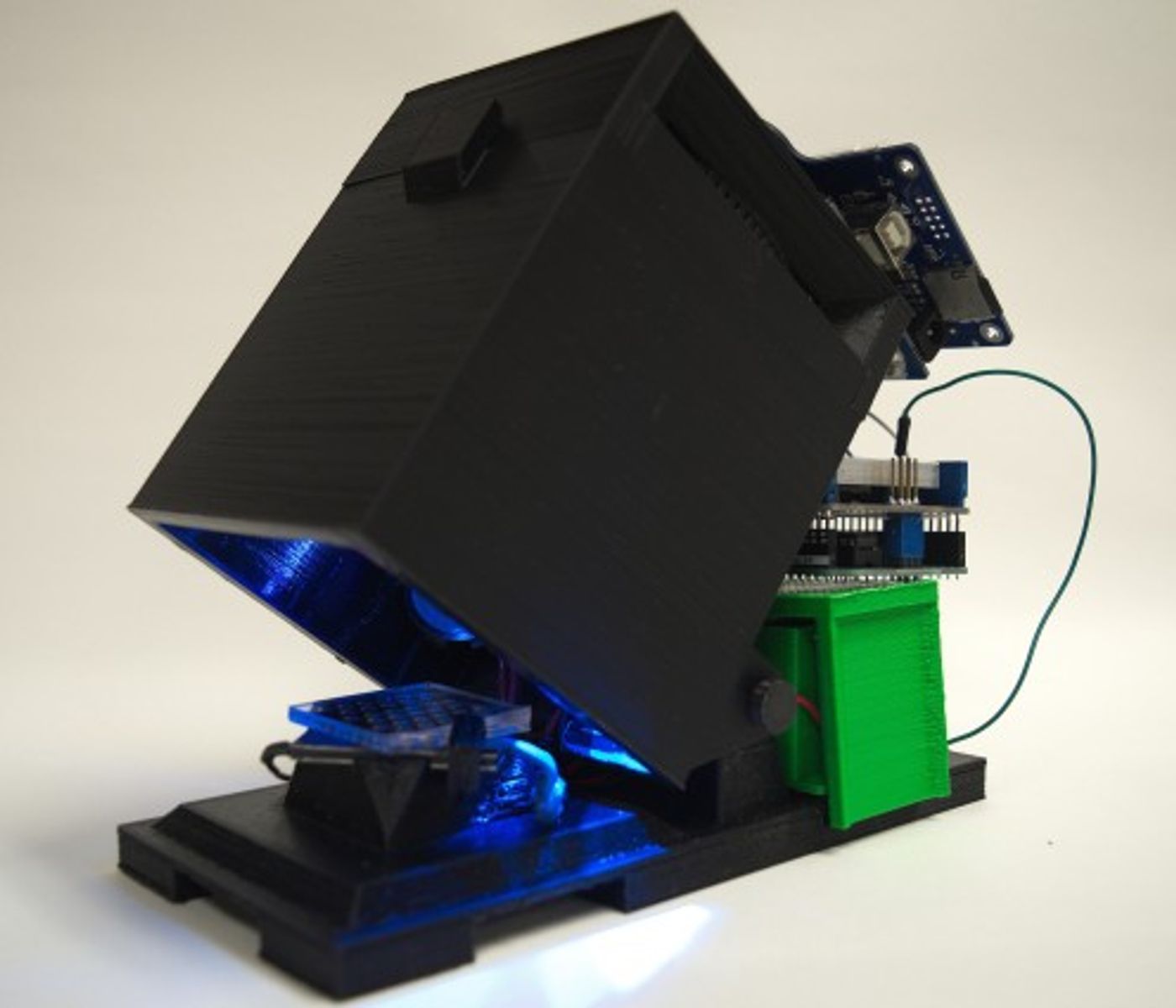
A research team that included scientists at the Lawrence Livermore National Laboratory (LLNL) and collaborators working at the University of Wisconsin, Madison, created a portable diagnostic kit that is also easy to use and low in cost. The kit is toughly the size of a shoebox, and it runs on 9 volt batteries that use a soft plastic reader, a B-chip or bacteria-detection chip, to perform a recombinase polymerase amplification (RPA) assay in around one hour (learn more about RPA in the video below). The chip utilizes wells carried on a disposable plastic assay cartridge that is then able to assess up to 16 different diseases at the same time. Sample solutions that have been degassed are self-pumped into an inlet and air is subsequently added to form a barrier that physically separates the reactions into the wells, preventing any cross-contamination.
"Making progress toward timely and accurate pathogen identification for an infection is the critical first step in effective patient care, especially in resource-limited environments, enabling proper usage of antibiotics to mitigate the growing emergence of antibiotic-resistance," explained LLNL principal investigator Tuan Nguyen.
The team has tested the kit against a variety of bacterial infections referred to collectively as ESKAPE (Enterococcus faecium, S. aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa and Enterobacter) organisms, so named because they can frequently elude the effects of many antibiotic therapies. Scientists opted to use RPA instead of polymerase chain reaction (PCR) assays in their tool.
"Isothermal DNA amplification methods represent an alternative to established techniques, such as real-time PCR, that require sophisticated equipment or complex experimental procedures," explained Nguyen. "RPA reactions are sensitive, specific and rapid and operate at constant low temperature that minimizes power consumption and simplifies heating and power handling."
Being able to quickly identify bacteria may also aid in the development of effective chemotherapy treatments. It may also improve industries that aim to maintain sanitary environments like the ones required in some types of manufacturing or food processing. This technology could also open up new avenues that improve quality control both by lowering costs and widening their scope.
"This demonstrated what great things can happen when chemists, biologists and bioinformatics teams come together," commented Tom Slezak, LLNL team leader of Pathogen and Bioinformatics for the Global Security Directorate. "We love to work on complex problems and translate what we do in biodefense to human health. It is a wonderful example of a national lab and academic collaboration."
Sources: Phys.org via LLNL, Applied and Environmental Microbiology