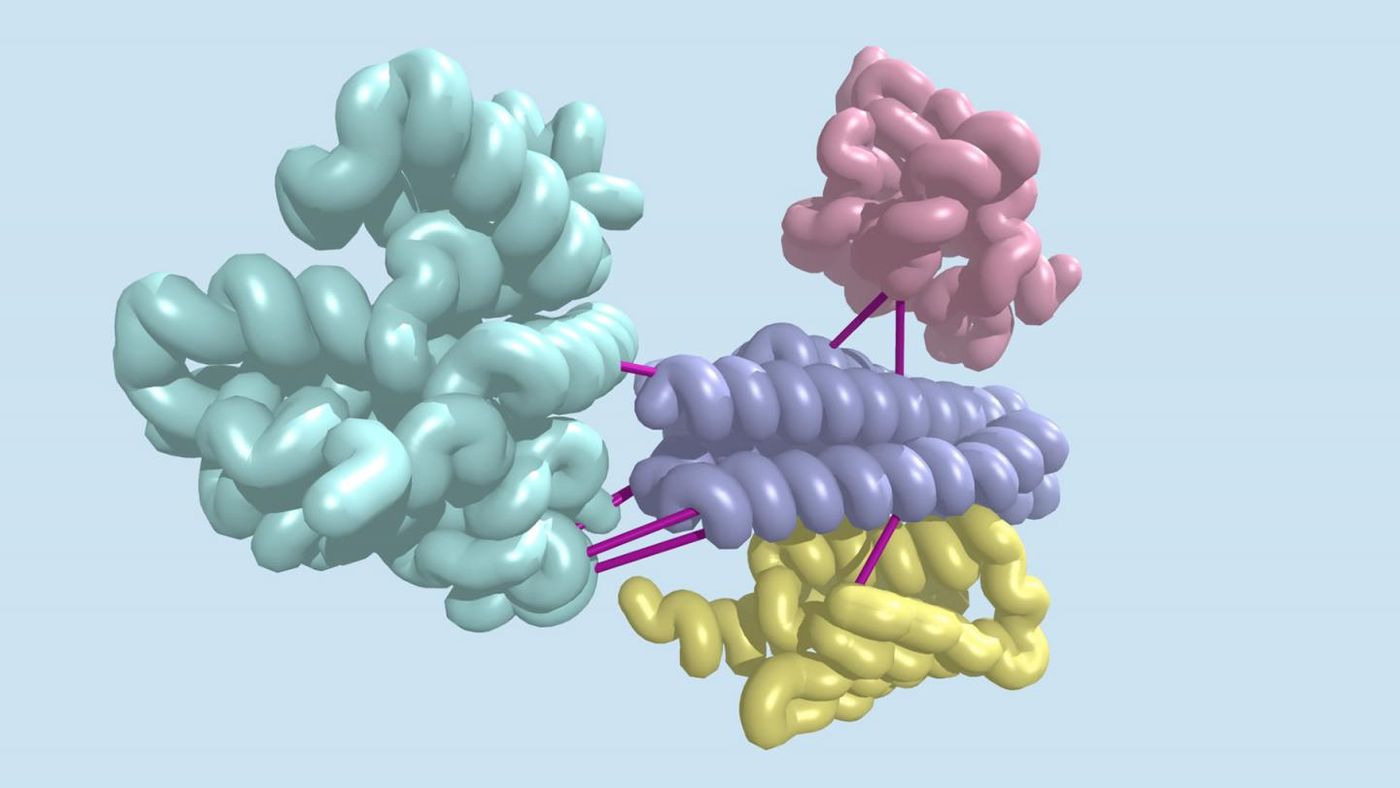
Revealing Protein Interactions by Studying the Genome
Proteins carry out the critical processes necessary for life. Having an understanding of biology requires revealing the relationships and interactions between proteins. While there are a variety of methods for finding those interactions, it can be a laborious, expensive, and time-consuming prospect. Now scientists have utilized advances in genetic sequencing technology to make it easier to identify protein interactions.
Reporting in Science, researchers at Harvard University and the University of Washington School of Medicine have found evolutionary signatures that bacterial gene pairs have in common. Using this approach, the scientists discovered hundreds of protein interactions that were unknown before this. Now the technique is being used to find protein interactions in humans.
"Protein-protein interactions are fundamental to biological function. It's remarkable that they can now be predicted en masse using the large amounts of genomic sequence data that have been generated in recent years," said the senior author of the study David Baker, professor of biochemistry at the University of Washington (UW) School of Medicine.
In this work, computational biologists used the co-evolution phenomenon, in which changes in one gene are linked to changes in another; it suggests that the two genes have an important connection. If two genes produce proteins that interact and one becomes mutated, causing its shape to change, the other gene might then evolve so that the protein it generates will continue to interact with its mutated partner, for example.
"Co-evolution has been useful for understanding how specific proteins interact, but we can now use it as a tool for discovery," said lead author Qian Cong, a postdoctoral fellow at the UW School of Medicine.
The investigators compared over 4,000 E. coli genes to 40,000 other bacterial genomes. Using statistics, they analyzed co-evolution among E. coli genes. The researchers identified 1,618 pairs of genes that probably co-evolved. The team compared their findings to known interactions; their method had better accuracy than other screening methods that are being developed.
It is unclear whether any of these predicted interactions were verified with traditional benchtop methods like yeast two-hybrid, but some may help explain some known biological observations. More work is also being planned.
"It is rare in biology for a software tool to make predictions that are promising enough to test, but that is exactly what's happening here," noted Cong. There are literally hundreds of follow-up experiments that could be performed in labs around the world."
The scientists also assessed protein interactions in the pathogenic bacterium Mycobacterium tuberculosis with this tool. They found 911 interactions, 95 percent of which were previously unknown. Some involve virulence and may help researchers create new therapeutics for tuberculosis.
"We are going to apply this tool to more pathogens, and the human genome," added Cong. "Our success will depend on how much work other scientists put into annotating which parts of the genome are genes and which parts are something else."
Sources: AAAS/Eurekalert! Via University of Washington, Science