Researchers Explore the Electricity-Conducting Power of Proteins
Enzymes are critical molecules that act as catalysts for many crucial reactions in a variety of cellular pathways. New work by researcher Stuart Lindsay and his team has identified a function of enzymes and potentially, all proteins; they can conduct electricity under certain conditions. Protein conduction is not a well-understood process, and this could help open up new approaches in medical devices and industrial production. One enzyme in particular may help advance rapid DNA sequencing technologies, for example.
"It is a way of plugging the amazing chemical diversity of enzymes directly into a computer," Lindsay explained. This research, which has been reported in ACS Nano, "opens the Pandora's box of looking at the function of any enzyme in a computer chip."
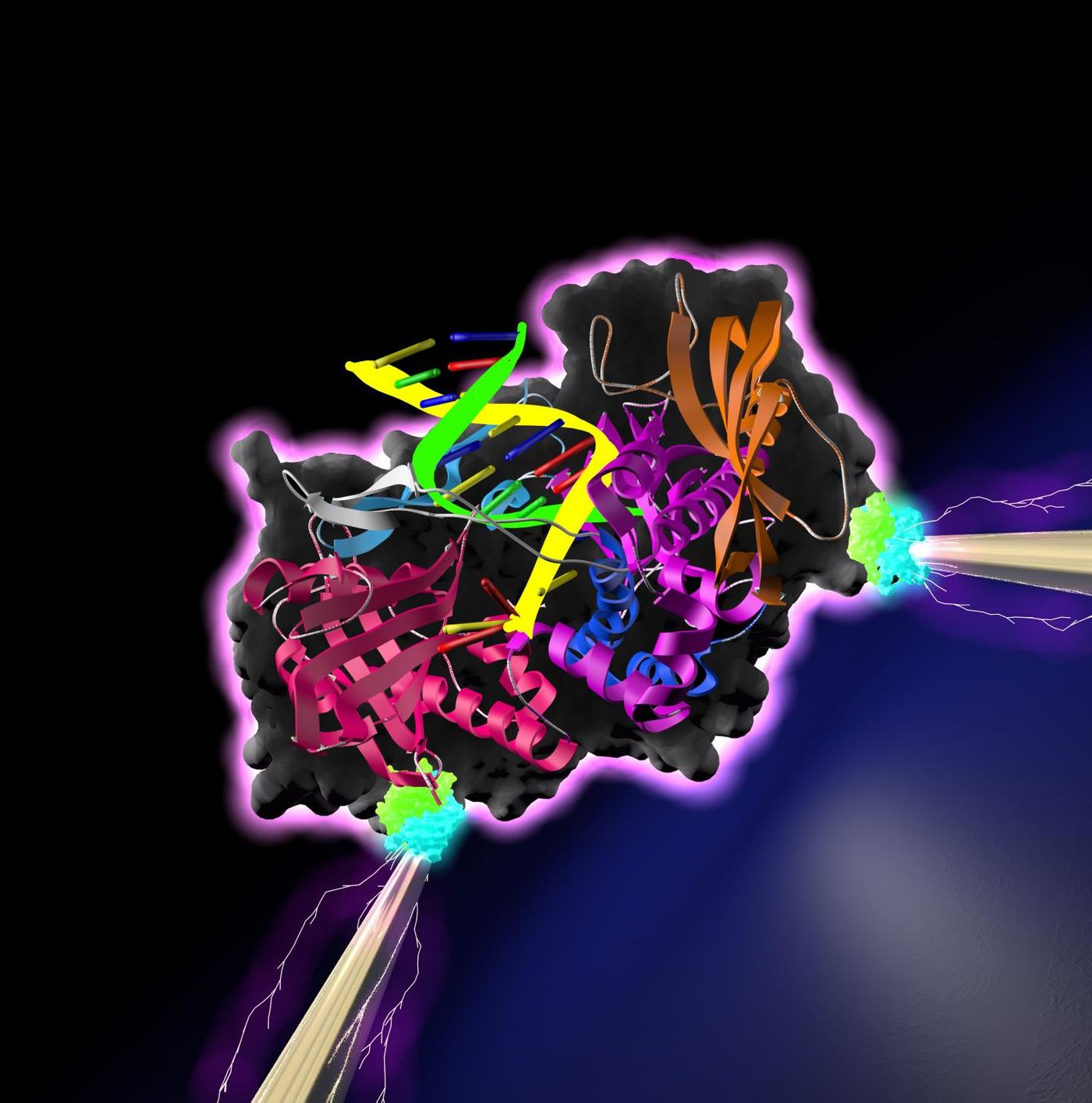
In this work, reported in ACS Nano, the researchers managed to attach an enzyme that repairs or synthesizes DNA called a DNA polymerase to a pair of electrodes. They measured spikes of current as the enzyme was bound to and released from target DNA bases. Their methodology shows that protein arrays can be mounted onto computer chips, where they could function as processors for various tasks.
"Enzymes are incredible molecules that carry out chemical reactions that just wouldn't happen otherwise," said Lindsay, who is the Director of the Biodesign Center for Single Molecule Biophysics at Arizona State University.
Researchers have known that proteins can insulate electrical flow, but their power as conductors has only recently been recognized. Now, they may be positioned in between insulators and conductors.
Previous work by Lindsay has shown that electricity can be conducted by a protein that is poised between two electrodes. The regions of protein that bind other molecules and exert its effects, its active sites, were hooked up to the electrodes. This new research linked a protein to electrodes using different binding sites, and the active sites could still function naturally.
The scientists linked the DNA polymerase to electrodes using the molecules biotin and streptavidin, which can act as linkers, and strong conductance signals were generated as the polymerase successively bound and released nucleotide bases.
"Wouldn't it be neat if you could put a pair of electrodes inside of polymerases because the polymerase grabs the DNA and chugs it through the junction. If you had a readout mechanism embedded in the polymerase, you've got the ideal sequencing machine," explained Lindsay.
The biotin molecule seems to be critical in generating conductive signals, and attaching the enzyme to electrodes is a painstaking process. In applying this technique, researchers have to be careful not to disrupt the active sites of the protein. For sequencing, conductance signals from each nucleotide base would have to be separated from background noise. Machine learning might help improve the technique. If the polymerases are isolated and incorporated into sealed chips that hold them into place, Lindsay suggested that these hurdles can be overcome.
This technique has the potential to speed up DNA sequencing to a breakneck pace - around one hundred nucleotides per second.
"If you put 10,000 molecules on a chip -- not a hard thing to do -- you'll sequence a whole genome in under an hour," Lindsay said. With current tools, an entire human genome takes about one day to sequence.
Sources: AAAS/Eurekalert! via Arizona State University, ACS Nano