L. monocytogenes is a bacteria that can be found anywhere in nature. It is a psychrotolerant pathogen, meaning, that it is able to survive and even grow at refrigeration temperatures. Outbreaks of listeriosis generally take place over a long period of time and are caused by refrigerated ready-to-eat foods such as milk, cheeses and deli meats. According to a recent report released by the Interagency Food Safety Analytics Collaboration (IFSAC) Project,
L. monocytogenes is considered a high priority pathogen due to the frequency and severity of the illness it causes as well as its susceptibility to targeted interventions. It is estimated to be the cause of 94 % and 15.9 % of hospitalizations and mortalities, respectively, linked to foodborne illness.
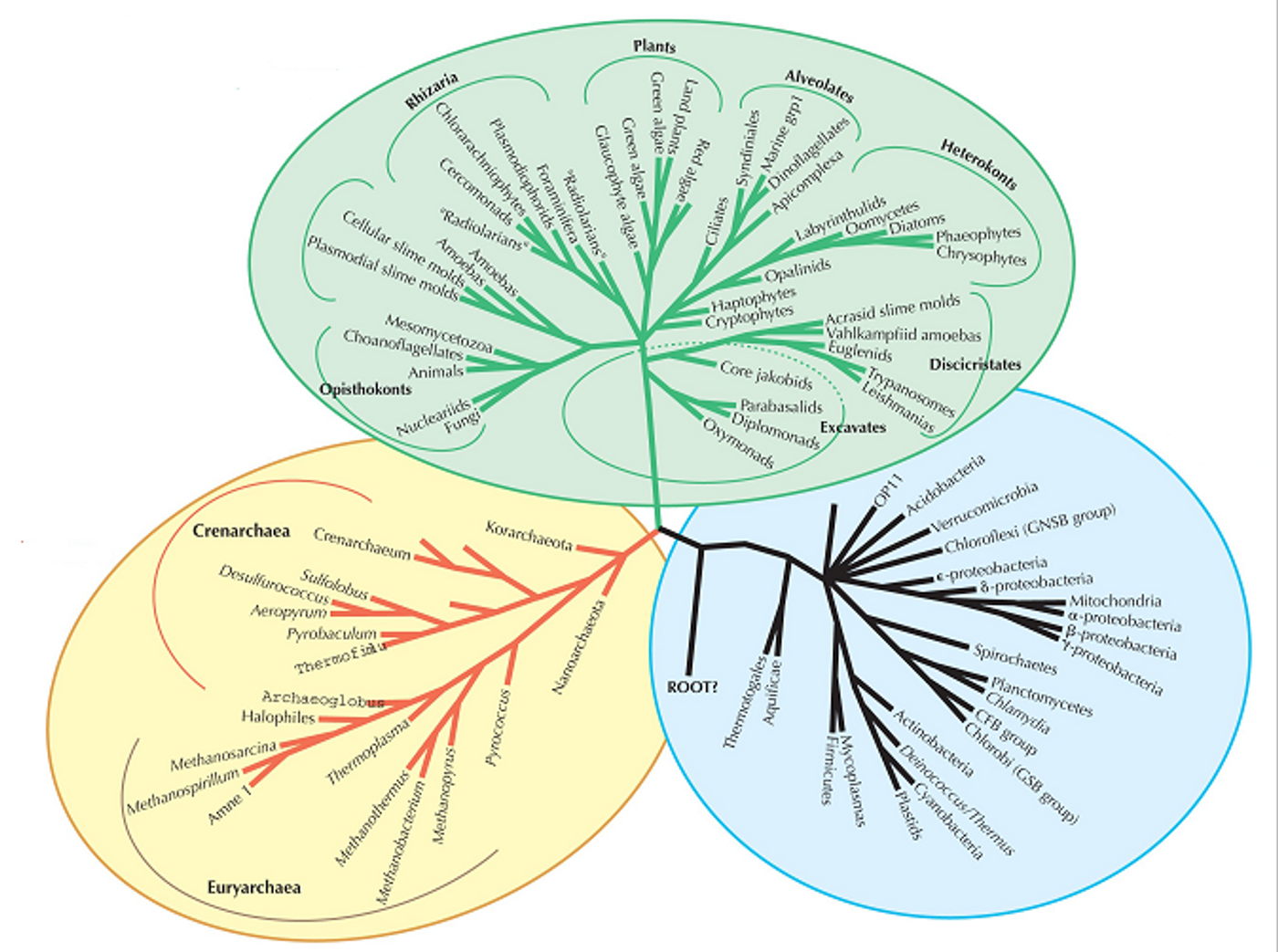
L. monocytogenes isolates divide into four distinct phylogenetic lineages. Isolates in lineage I are responsible for the majority of foodborne outbreaks. Isolates in lineage II have been implicated in large epidemics and are the cause of sporadic cases of foodborne illness. Lineage III and IV isolates are rarely associated with disease in humans.
Scientists from North Dakota State University used a molecular typing technique to investigate the genetic diversity of 124
L. monocytogenes isolates from various sources in order to determine if there was any correlation between the ability of isolates to cause disease and their genetic makeup. They focused on isolates from lineages I and II since they are most associated with disease in humans. Isolates were inoculated into a growth medium that was limited in nutrients to test the ability of each isolates to grow in an environment not ideal for growth, such as a food processing environment. They also used multi-locus sequence typing (MLST) to determine the genetic characterization of each isolate.
The isolates had variable growth rates in the nutrient-limited medium however; scientists were unable to determine if there was a link between growth rate and a specific lineage or serotype. Isolates in lineage I had significantly lower growth rates compared to those in lineage II. Genetic characterization using (MLST) revealed that the 124 isolates represented 81 different sequence types and 33 different clonal complexes, indicating the diversity of the isolates used in this study. The authors were unable to make any correlations between serotype or source of the isolates (human, food, environment or animal) and growth rate. However; authors concluded that the ability if lineage II isolates to grow faster than lineage I in nutrient-limited medium may facilitate their ability to persist in non-host environments leading to a higher risk of food contamination and transmission of the pathogen through food to humans. This study also highlights the importance of molecular subtyping in evolutionary analysis in pathogens to facilitate surveillance, detection and control of pathogens.
Source:
Foodborne Pathogens and Disease;
IFSAC;
Emerging Infectious Diseases; LabRoots; Genetic Diversity in Microorganisms