‘Junk DNA' Exposed - Transient Gene Expression Can Now Be Studied
Long regions of seemingly unimportant strands of DNA were named ‘junk DNA’ and long thought to be unimportant or at least, totally misunderstood. That view is shifting, however, as more and more regulatory signals are found to be in those areas; mutations in them can contribute to human disease.

Typically, a gene in DNA will encode for a protein. Those genes, the corresponding RNA, and their protein products can be analyzed and manipulated in many ways as part of basic molecular biology. But not all genes are expressed in every place in the body at all time. Controlling that expression is vital to the proper function of cells, which makes the regulatory portions of DNA very important. The DNA that ends up getting expressed in cells as RNA is called the transcriptome.
"Regulatory DNA regions are essential for development in humans, tissue preservation, and the immune response, among others," explained Professor Patrick Cramer, a director at the Max Planck Institute for Biophysical Chemistry. "Furthermore, they play an important role in various diseases. For example, patients suffering from cancer or cardiovascular conditions show many mutations in exactly those DNA regions," he said.
Identifying and studying those governing regions is difficult though, because when DNA that does the regulating is active, it’s made into short-lived RNA transcripts that are used and then quickly degraded. But those fleeting transcripts that are so challenging to study are crucial activators of genes and play a prominent role in gene expression.
Scientists have now developed a way to capture those transcripts so they can be analyzed in depth. Researchers collaborating and working with Dr. Cramer have created a precise way to identify and capture RNA molecules, even those with a brief, transitory existence. The work was published in Science. They’ve called the new technique the TT-Seq (transient transcriptome sequencing) method, and it can gather and sequence all RNA segments that are made in cells over a period of 5 minutes – an insufficient amount of time to degrade even very transient RNA.
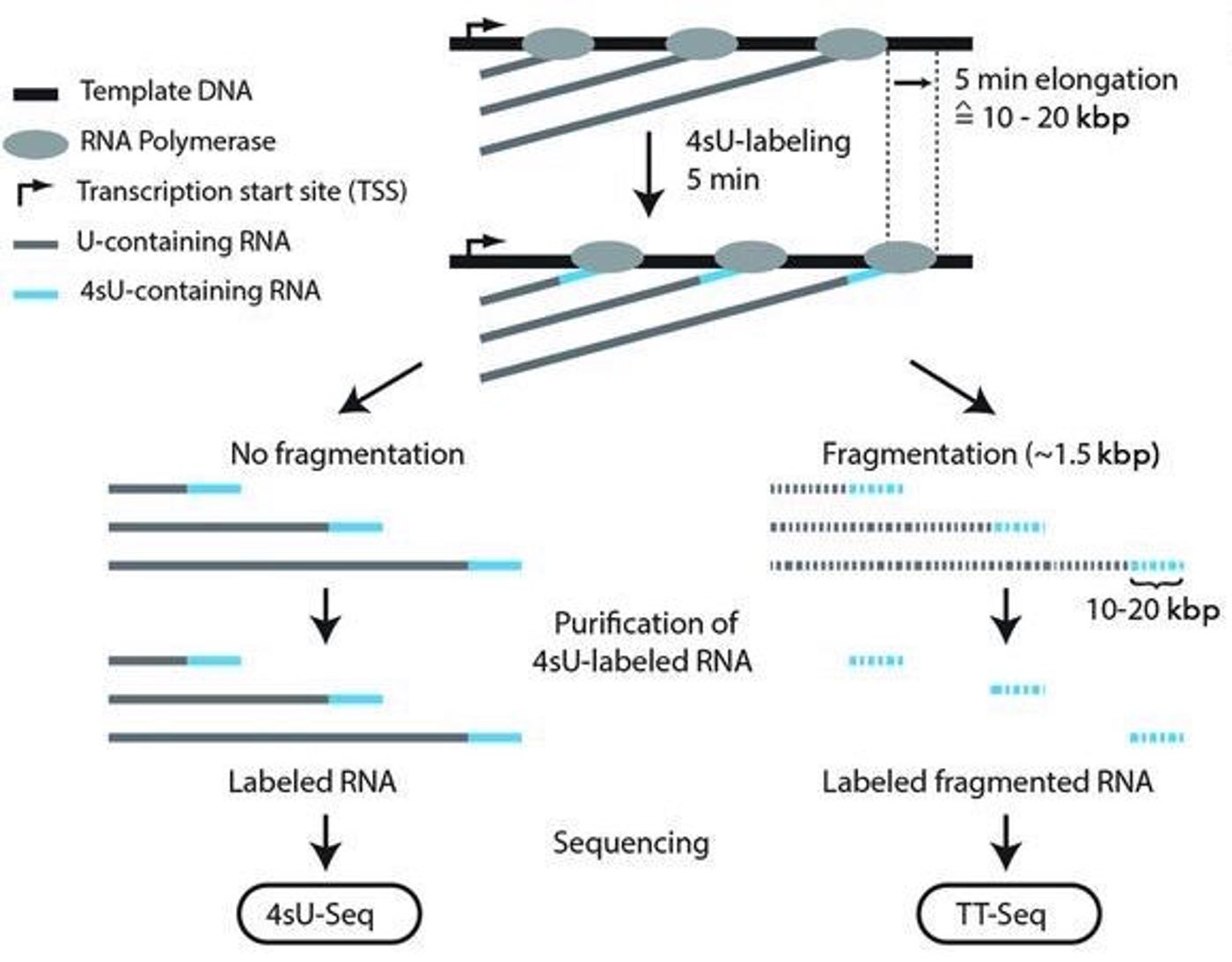
They use a molecule, 4-thiouridine (4sU), that gets incorporated into the RNA as it is made for a kind of tag, and then isolate whatever contains that tagging molecule for sequencing.
"The RNA molecules we caught with the TT-Seq method provide a snapshot of all DNA regions that were active in the cell at a certain time – the genes as well as the regulatory regions between genes that were so hard to find until now," Cramer said. "With TT-Seq we now have a suitable tool to learn more about how genes are controlled in different cell types and how gene regulatory programs work," added co-author Julien Gagneur, Professor of Computational Biology at the Technical University of Munich.
To learn more about the transcriptome, check out the video below.
Sources: Science, Technical University of Munich