Glucocorticoids are steroidal hormones that signal through the glucocorticoid receptor in complex ways throughout the body. Researchers have known for some time that the glucocorticoid receptor is able to bind the genome in many places, over 10,000, in fact. However, the receptor is only responsible for regulating the expression of a few hundred genes. Why there are so many places where it would apparently act to start protein production with so few in use was a mystery. Not only are there many receptor binding sites in the genome, the sites are not distributed evenly or randomly. Those sites tend to be clustered around certain parts of the genome. Scientists decided to investigate by using relatively new high-throughput gene analysis techniques.
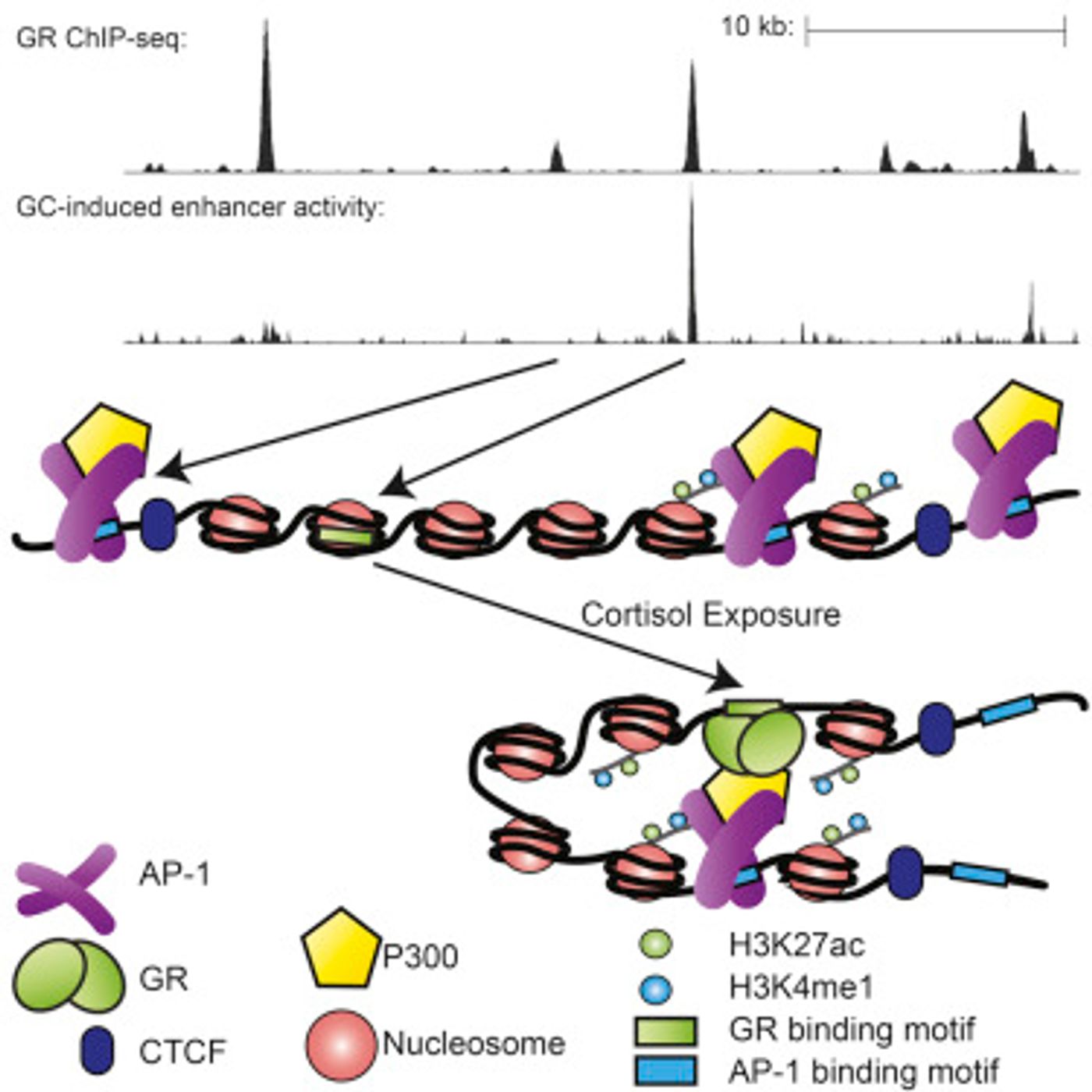
The research team, led by Tim Reddy, an assistant professor of biostatistics and bioinformatics at Duke University, focused on the cortisol production pathway, a hormone signaling system that is a critical part of the human stress response. The team utilized a cell line from human lungs; "we did millions of experiments over tens of thousands of sites on the genome," Reddy said. "We were really surprised to find that only about 13 percent of these binding sites seemed to do anything when you chopped them out of the genome and looked at them alone."
The researchers termed that 13 percent the “direct-response sites.” They learned that those sites were not particularly powerful when it came to signaling. Reddy was quick to add that the remaining 87 percent weren’t just useless leftovers either. Instead, the two seem to work in collaboration to amplify a hormone signal. They
reported in Cell that when the inactive sites are brought together with a direct-repsonse site, there is a 100-fold increase in the signal that goes to the gene.
"Our hope is that learning more about how these clusters coordinate to turn on genes in different cells will help us design new drugs that only act on the cells that we want," said the lead author on the study, Duke Cell Biology graduate student Chris Vockley.
While exercising with his wife, Reddy devised an analogy for this organization of binding sites: the cell is like a house while the direct-action binding sites act as the electrical outlets in various rooms.
"These are the places in the genome that directly respond to a hormone," Reddy said. "The appliances in the house end up clustered around these outlets because the cords are only so long, just as we see indirect sites clustered around direct sites in the genome."
And if one family moves out of the house, and another moves in -- representing a cell of a different tissue type -- the appliances may be plugged in with a different configuration. "The outlets don't change, but the appliances do."
"It really seems like at the core of these clusters, there might be one or a small number of sites that are really the main 'electrical outlet' that everything can plug into and everything else can kind of loop in," Reddy said.
"The most important thing to me is that this paper shows that glucocorticoid receptors don't -- at least in this system -- act to directly repress gene expression," Guertin continued, controverting a widely-held belief.
"It doesn't surprise me that it's more complicated than some people hoped it would be, explained Mike Guertin, an Assistant Professor Biochemistry and Molecular Genetics at the University of Virginia, who was not involved in this study. "The field is pretty nascent. We've only been able to look at whole-genome questions like this since about 2007. I think we have a lot of opportunities to design clever, hypothesis-driven experiments like this, instead of just generating data."
Reddy is ready for the challenge. "If the genome is no longer a bunch of independent sites that kind of add up to some response, but instead has interactions between all these sites, we either have to fundamentally change how we're modeling the whole genome -- which is a huge thing - or it means that we need to start thinking about these groups of sites, understanding why genes are regulated and why disruption of that regulation can cause disease.”
To learn more about glucocorticoids, their receptors, and how they influence genetic expression in cells, view the video above.
Sources:
AAAS/Eurekalert! via
Duke,
Cell