In the natural world, genes don’t always operate independently. They can function as part of a set of genes and can be regulated at many levels, including by one another in feedback mechanisms that are both positive and negative in nature. Those relationships can in turn be influenced by changes in the genome, or mutations. A deep understanding of how genes work and how they contribute to disease requires elucidating how they interact to coordinate the functions of life.
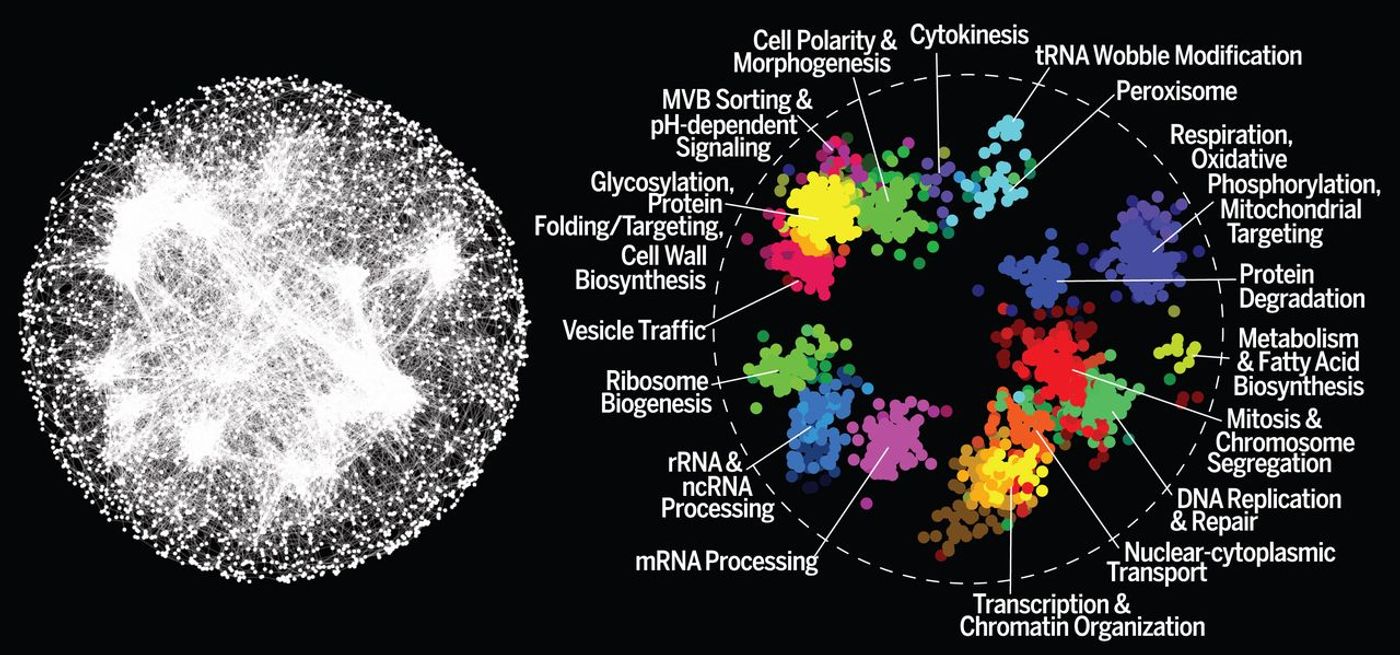
A landmark new study led by University of Toronto's Donnelly Centre Professors Brenda Andrews and Charles Boone, and Professor Chad Myers of the University of Minnesota-Twin Cities has generated the first map illustrating the genetic interaction profile of a cell, shown above. This incredible work, which took 15 years to finish, unveils a new tool for the exploration of the genetic impact on disease with an eye toward the development of more precise therapeutics. The
report has been published in the journal Science.
"We've created a reference guide for how to chart genetic interactions in a cell," explained one of the researchers who spearheaded the study, Michael Costanzo, who is a research associate in the Boone lab. "We can now tell what kind of properties to look for in searching for highly connected genes in human genetic networks with the potential to impact genetic diseases."
Yeast cells have functioned as a model for genetic studies for many years. They have 6000 genes and many are also found in humans. About a decade ago, a consortium of researchers completed a massive undertaking to delete each yeast gene individually and observe the effects. They found that only about 20 percent of the genes were required for the survival of the organism. Last year, human cells underwent a similar analysis with a similar result; only a fraction of human genes are required.
Those findings have suggested that genes are protected or buffered from mutations and environmental challenges. To get at that question, investigators knew they would have to test millions of gene pairs to see how cells fare when they lose more than one gene at a time.
To search for those essential gene pairs, Andrews, Boone and Myers led the work to delete genes in combination. That meant a custom-built robot had to be made in order to analyze the 18 million various combinations. Results made it clear that many genes have one or more backups for ensure functionality if they are lost. That’s led the scientists to conclude that instead of focusing on single genes in human disease, gene pairs should be under study instead.
The new map will help the understanding of how genes interact in cells, and it’s hopes the map will expand to include a variety of cell types. When used in combination with whole genome studies and personalized medicine, gene combinations that contribute to disease should become more obvious.
"Without our many years of genetic network analysis with yeast, you wouldn't have known the extent to which genetic interactions drive cellular life or how to begin mapping a global genetic network in human cells," said Boone, who also holds a Canada Research Chair in Proteomics, Bioinformatics and Functional Genomics. “We have tested the method to completion in a model system to provide the proof of principle for how to approach this problem in human cells. There's no doubt it will work and generate a wealth of new information."
If you would like to know more about Professor Andrews' research, you can watch a talk she gave (above) about the using genomics to understand basic biology, with the help of yeast, at the National Insitutes of Health.
Source:
Phys.org via
University of Toronto,
Science