Labroots 2021 Clinical Research & Diagnostics Virtual Event's Winning Poster
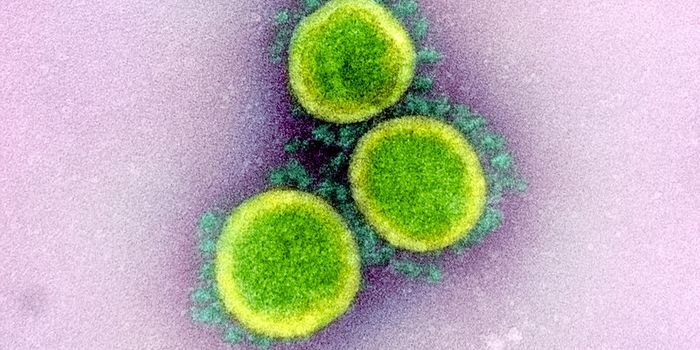
The SARS-CoV-2 virus that emerged in December 2019 causes the disease COVID-19, a pandemic that has killed over 5 million people worldwide. While genomic tools have enabled researchers to assess the sequence of the viral genome to create several vaccines, treatments have been more elusive. Though a few have now been developed, they are expensive and have to be taken soon after the onset of infection to be effective; more therapeutics are desperately needed to treat cases caused by existing strains of the coronavirus, as well as others that may emerge.
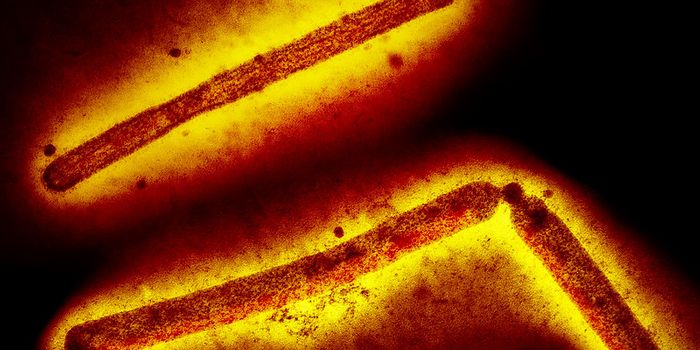
The SARS-CoV-2 virus is known to use a spike protein to latch onto a receptor on the surface of human nasal epithelial cells to cause infection. The viral spike protein is a glycoprotein that can bind to at least one receptor that we know of called ACE2 (angiotensin converting enzyme 2).
The winners of best poster at the Labroots 2021 Clinical Research & Diagnostics Virtual Event were poster author Yugmee Gidiya, and mentor Dr. Rahna. K. Rathnan of the JSS Private School in Dubai. They examined studies by other research groups that have investigated the phylogenetic analysis and homology modeling of the SARS-CoV-2 virus. Phylogenetic tools can help scientists classify and understand the biology and evolution of an organism and its genes. In this case, phylogenetics showed researchers the relationships between the strains of SARS-CoV-2. In homology modeling, scientists can assess the predicted three-dimensional structure of a protein, whether it be the viral spike protein or other proteins produced by the SARS-CoV-2 virus, such as viral proteases. Computational tools can be used to investigate the interactions, including the thermodynamic characteristics, of the viral spike protein and the human ACE2 receptor, or other viral proteins and their human binding partners.
The binding that occurs between the amino acid residues of two different proteins can be extremely complex. But if the binding free energy between a viral protein and its binding partner can be accurately calculated, it may be possible to identify drugs that can disrupt the interaction between those two proteins to prevent SARS-CoV-2 infection or treat one after it's happened.
This poster cited a paper reported in Chemical Physics in early 2021 that suggested that five molecules called protease inhibitors can bind to a COVID-19 protein that functions as an enzyme called a protease, and those inhibitors could be useful in preventing or treating SARS-CoV-2 infection. One protease inhibitor in particular called lopinavir was theorized to have the greatest potential against COVID-19.
This poster included strains that existed prior to the emergence of the Omicron variant, and up to that point, the virus appeared to be mutating slowly; only a few base pairs had changed among the different strains. That trend may have changed with the emergence of Omicron, a variant that did not evolve from the Delta variant but instead came from an earlier version. Omicron has at least 50 mutations, which are different from those seen in the Delta variant.