From the Sanger Institute/ EMBL-EBI Single Cell Genomics Centre, scientists say it is now possible to assay both DNA methylation and RNA of the same single cell at the same time.
While the genome remains stable, the transcriptome is constantly changing. The transcriptome is made up of messenger RNA molecules that vary in production based on environmental factors and the stages of development (
Nature). On the other hand, the epigenome is the collection of chemicals that control what proteins the genome produces (
Genome.gov). In a recent study published in
Nature Methods, scientists have put finishing touches on a new protocol to study the interaction between the transcriptome and the epigenome simultaneously.
“To understand development, it is really important that we pin these relationships down, and get them right,” said Wolf Reik of the Babraham Institute and Wellcome Trust Sanger Institute.
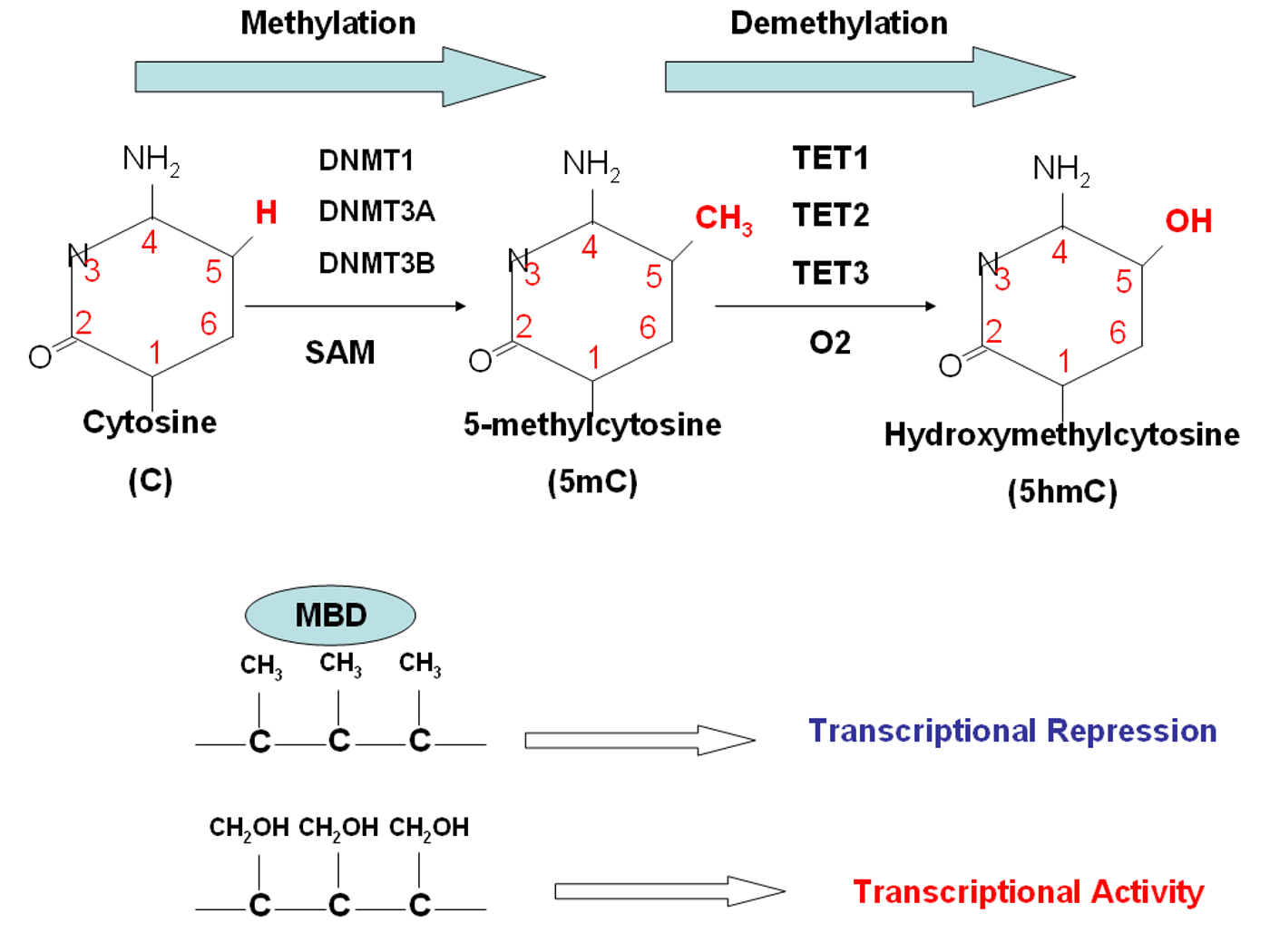
Using mouse embryonic stem cells (ESCs), the research team used “two techniques in parallel,” one for looking at the transcriptome and the other for looking at epigenetic changes from DNA methylation. ESCs never fully commit to a differentiated state, so the researchers could visualize transcriptome changes from DNA methylation as the cells switched back and forth between different gene expression states. The team was able to gain information about several thousand genes.
Using this dual approach, scientists can gain novel insight into how ESCs maintain pluripotency and how the regulate cell differentiation. Additionally, the research team hopes to make connections between gene expression and epigenetic changes for normal development as well as deviations that could explain cancer pathology and aging.
“The result is an optimized approach that maximizes the amount of biological information that can be obtained from a single cell,” said Thierry Voet of the Wellcome Trust Sanger Institute and Katholieke Universiteit Leuven, Belgium.
Source:
European Bioinformatics Institute EMBL-EBI