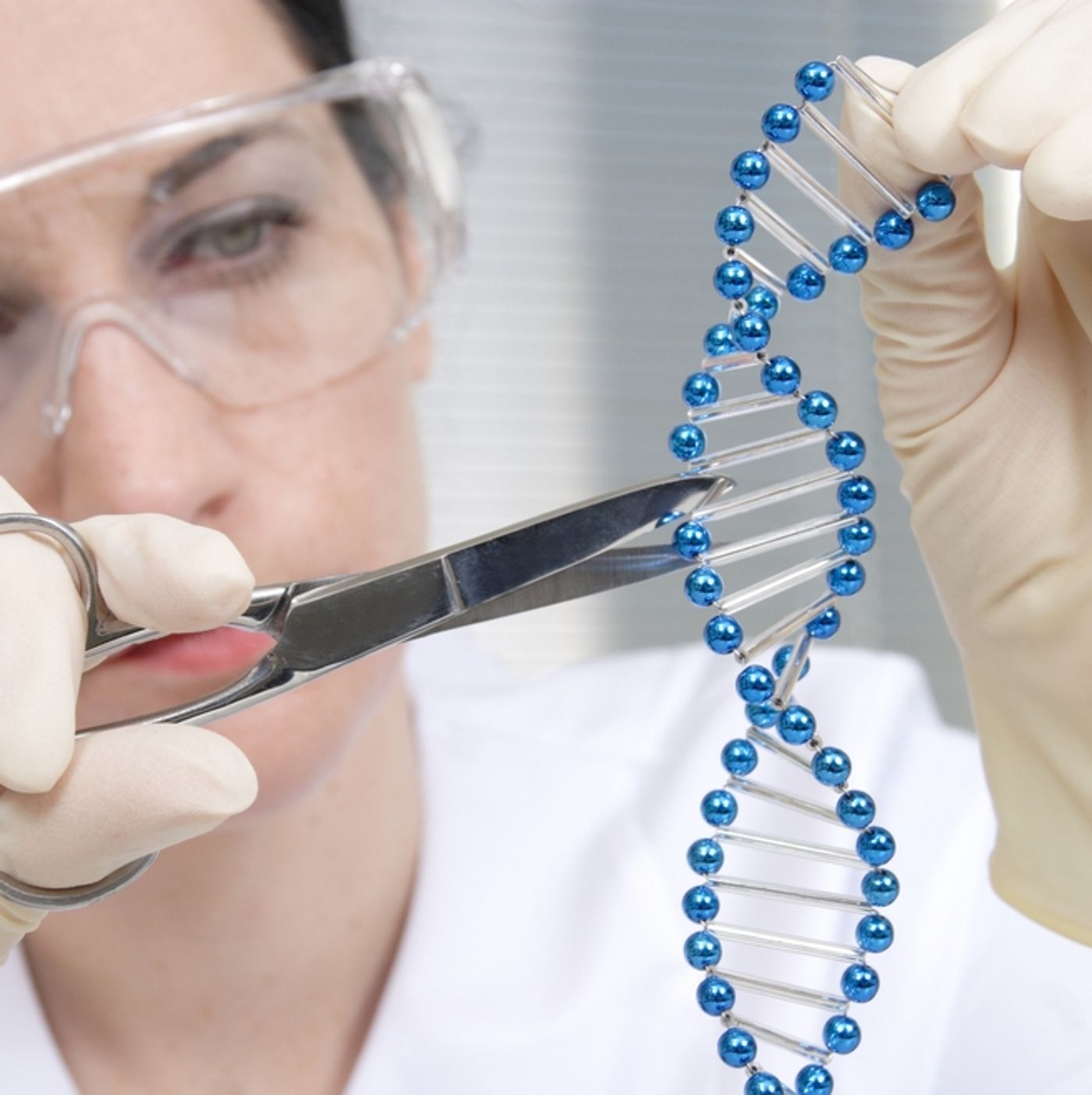
CRISPR-Cas9 is the discovery that took the scientific community by storm, the technology of the century. In a new study published in two
Nature Biotechnology papers, researchers introduce a new player in CRISPR genome editing: Cpf1.
“Clustered Regularly Interspaced Short Palindromic Repeats” is the name for an adaptive immunity mechanism found in bacteria in the 1980s then applied almost a decade ago by Rudolphe Barrangou, PhD. Since then scientists have often used an endonuclease called Cas9 that can be programmable by small RNAs to cleave target DNA. Researchers from the Institute for Basic Science (IBS) Center for Genome Editing now offer Cpf1 as an alternative endonuclease to Cas9.
What makes Cpf1 a better option than Cas9? The researchers from IBS offer a few convincing points.
- Cpf1 requires only a single guiding RNA
- Cpf1 recognizes thymidine-rich DNA sequences, whereas Cas9 recognizes guanosine-rich sequences
- More efficient genome sequencing
- More diverse mouse models of mutation
To demonstrate how Cpf1 endonuclease finds its targets, the researchers conducted a test called “digenome-seq,” a procedure that can show how successful or unsuccessful a CRISPR endonuclease is at targeting the correct sequences for cleaving.
They used two Cpf1 family proteins to compare with Cas9: AsCpf1 and LbCpf, two family members of Cpf1 that are known to be the most efficient. From computational analysis, they could tell that Cpf1 endonucleases quickly degraded after they successfully cleaved a target site, eliminating the opportunity for unnecessary cleavage afterward.
“Thereby reducing off-target effects without sacrificing on-target effects," said KIM Jin-Soo, the corresponding author of the two papers and Director of the IBS Center.
Compared to Cas9, Cpf1 proved to be highly specific, cleaving fewer off-target sites than Cas9. For the majority of in vitro cleavage sites, the off-target cleavage events for Cpf1 were less than 0.1 percent.
Cpf1 RNP-mediated mutations were also introduced into mouse embryos, targeting both Foxn1, a transcription factor that regulates the immune system, and tyrosinase, an enzyme that prompts the production of melanin. The mutant embryos were than transplanted into surrogate mothers, who gave birth to mice with intended mutations in Foxn1 and tyrosinase. Following whole-genome sequencing from genetic material from these mice, researchers observed zero off-target mutations in both mutants as compared to their normal, wildtype siblings.
The mutant mice showed their mutations phenotypically, as hairless and white-haired for the Foxn1 and tyrosinase mutants respectively. Cpf1 RNP in these mutants had knocked out intended genetic functions, 64 percent of Foxn1 and 33 percent of tyrosinase.
Scientists will continue to tinker with CRISPR genome editing technology, and this discovery is a great step forward in its improvement. Researchers anticipate the application of CRISPR-Cpf1 to be diverse and promising, from non-toxic cancer drugs to livestock and agricultural products to stem cell treatments.
Source:
Institute for Basic Science,
Nature Methods,
New England BioLabs