A Molecular On/Off Switch for CRISPR
TSRI Scientists Reveal How Viruses Disable Bacterial Immune Systems
LA JOLLA, CA – March 28, 2017 – Picture bacteria and viruses locked in an arms race.
For many bacteria, one line of defense against viral infection is a sophisticated RNA-guided “immune system” called CRISPR-Cas. At the center of this system is a surveillance complex that recognizes viral DNA and triggers its destruction. However, viruses can strike back and disable this surveillance complex using “anti-CRISPR” proteins, though no one has figured out exactly how these anti-CRISPRs work—until now.
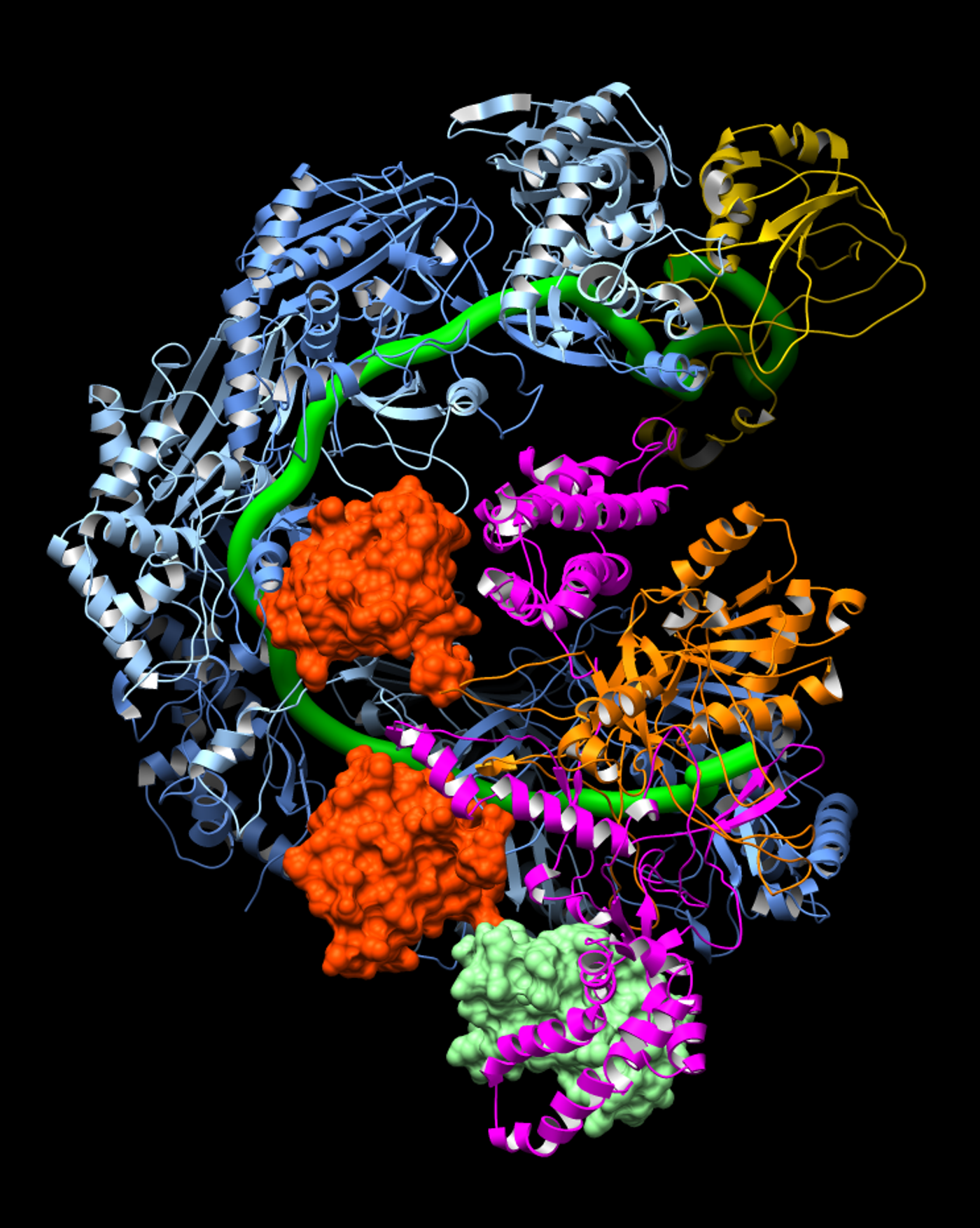
For the first time, researchers have solved the structure of viral anti-CRISPR proteins attached to a bacterial CRISPR surveillance complex, revealing precisely how viruses incapacitate the bacterial defense system. The research team, co-led by biologist Gabriel C. Lander of The Scripps Research Institute (TSRI), discovered that anti-CRISPR proteins work by locking down CRISPR’s ability to identify and attack the viral genome. One anti-CRISPR protein even “mimics” DNA to throw the CRISPR-guided detection machine off its trail.
“It’s amazing what these systems do to one-up each other,” said Lander. “It all comes back to this evolutionary arms race.”
The new research, co-led by Blake Wiedenheft of Montana State University, was published recently in the journal Cell.
If CRISPR complexes sound familiar, that’s because they are at the forefront in a new wave of genome-editing technologies. CRISPR (pronounced “crisper”) stands for “clustered regularly interspaced short palindromic repeats.” Scientists have discovered that they can take advantage of CRISPR’s natural ability to degrade sections of viral RNA and use CRISPR systems to remove unwanted genes from nearly any organism.
“Although CRISPR-Cas9 is the ‘celebrity’ CRISPR system, there are 19 different types of CRISPR systems, each of which may have unique advantages for genetic engineering. They are a massive, untapped resource,” said Lander. “The more we learn about the structures of these systems, the more we can take advantage of them as genome-editing tools.”
Using a high-resolution imaging technique called cryo-electron microscopy, the researchers discovered three important aspects of CRISPR and anti-CRISPR systems.
First, the researchers saw exactly how the CRISPR surveillance complex analyzes a virus’s genetic material to see where it should attack. Proteins within the complex wrap around the CRISPR RNA like a grasping hand, exposing specific sections of bacterial RNA. These sections of RNA scan viral DNA, looking for genetic sequences they recognize.
“This system can quickly read through massive lengths of DNA and accurately hit its target,” said Lander. If the CRISPR complex identifies a viral DNA target, the surveillance machine recruits other molecules to destroy the virus’s genome.
Next, the researchers analyzed how viral anti-CRISPR proteins paralyze the surveillance complex. They found that one type of anti-CRISPR protein covers up the exposed section of CRISPR RNA, thereby preventing the CRISPR system from scanning the viral DNA.
“These anti-CRISPR proteins keep the bacteria from recognizing the viral DNA,” Lander explained. He called these anti-CRISPR proteins “exceptionally clever” because they appear to have evolved to target a crucial piece of the CRISPR machinery. If bacteria were to mutate this machinery to avoid viral attacks, the CRISPR system would cease to function. “CRISPR systems cannot escape from these anti-CRISPR proteins without completely changing the mechanism they use to recognize DNA,” he said.
Another anti-CRISPR protein uses a different trick. Based on its location and negative charge, the researchers believe this anti-CRISPR protein acts as a DNA mimic, fooling CRISPR into binding this immobilizing protein, rather than an invading viral DNA.
“These findings are important because we knew that anti-CRISPR proteins were blocking bacterial defenses, but we had no idea how,” said Lander.
The researchers believe this new understanding of anti-CRISPR proteins may eventually lead to more sophisticated and efficient tools for gene editing. Perhaps anti-CRISPR proteins can be used in CRISPR systems to swoop in to block gene editing—or researchers could degrade anti-CRISPR proteins to trigger gene editing. “That might work as an on-off switch for CRISPR,” Lander said.
In addition to Lander and Wiedenheft, the study, “Structure Reveals Mechanisms of Viral Suppressors that Intercept a CRISPR RNA-Guided Surveillance Complex,” was led by first authors Saikat Chowdhury of TSRI and Joshua Carter of Montana State University. Other authors of the study were MaryClare F. Rollins, Sarah M. Golden, Ryan N. Jackson and Connor Hoffmann of Montana State University; Lyn’Al Nosaka of TSRI; Joseph Bondy-Denomy of the University of California, San Francisco; Karen L. Maxwell and Alan R. Davidson of the University of Toronto, Toronto, CA; and Elizabeth R. Fischer of the National Institutes of Health’s National Institute of Allergy and Infectious Diseases.
The study was supported by the National Institutes of Health (grants P20GM103500, P30GM110732-03, R01GM110270, Q1 R01GM108888, P20GM103474, F32 GM108436, DP2EB020402 and DP5-OD021344); the National Institutes of Health Intramural Research Program; the Canadian Institutes of Health Research (grant MOP-130482 and MOP-136845); the University of California, San Francisco, Program for Breakthrough Biomedical Research, funded in part by the Sandler Foundation; the National Science Foundation EPSCoR (grant EPS-110134); the M.J. Murdock Charitable Trust; and a young investigator award from Amgen. Lander is also supported as a Searle Scholar and Pew Scholar.
This article was originally published on Scripps.edu.