From Peter Parker’s fateful spider bite to Arthur pulling the sword from the stone at the beginning of his reign—everyone likes to know a hero’s origin story.
In this case, our heroes are potent antibodies from the body’s immune system that bind to and neutralize HIV—and the story could lead to a vaccine against AIDS.
In a new study, a team led by scientists at The Scripps Research Institute (TSRI) tracked how a family of these HIV-fighting antibodies develops over time. The research shows how a future vaccine might trigger the immune system to produce these antibodies more effectively.
“If you could produce these antibodies through vaccination, that would be a really fantastic start to preventing HIV,” said Ian Wilson, Hansen Professor of Structural Biology and chair of the Department of Integrative Structural and Computational Biology at TSRI.
The research was published this month, in the journal
Immunity.
Antibodies in Training
At boot camp, it takes time for soldiers to hone their response against a specific threat. Similarly, it takes time for immune system cells to identify a particular virus’s weak points and enlist antibodies that mutate, tweaking their structures to better bind to and neutralize the virus.
In the case of one family of HIV-targeting antibodies known as PGT121, it takes around two years for antibodies to mature enough to fight HIV. While this delay is too long to be effective in preventing infection, the existence of these antibodies inspired scientists to try to find ways to harness their power in a vaccine.
In the new study, scientists looked at the early stages of development in the PGT121 family, specifically in the PGT122 and PGT124 branches. These antibodies caught scientists’ attention because they are “broadly neutralizing,” meaning they can stop many strains of the rapidly mutating virus. They also have the special ability to target a piece of viral machinery called the HIV Envelope glycoprotein, which enables the virus to bind to and penetrate host cells.
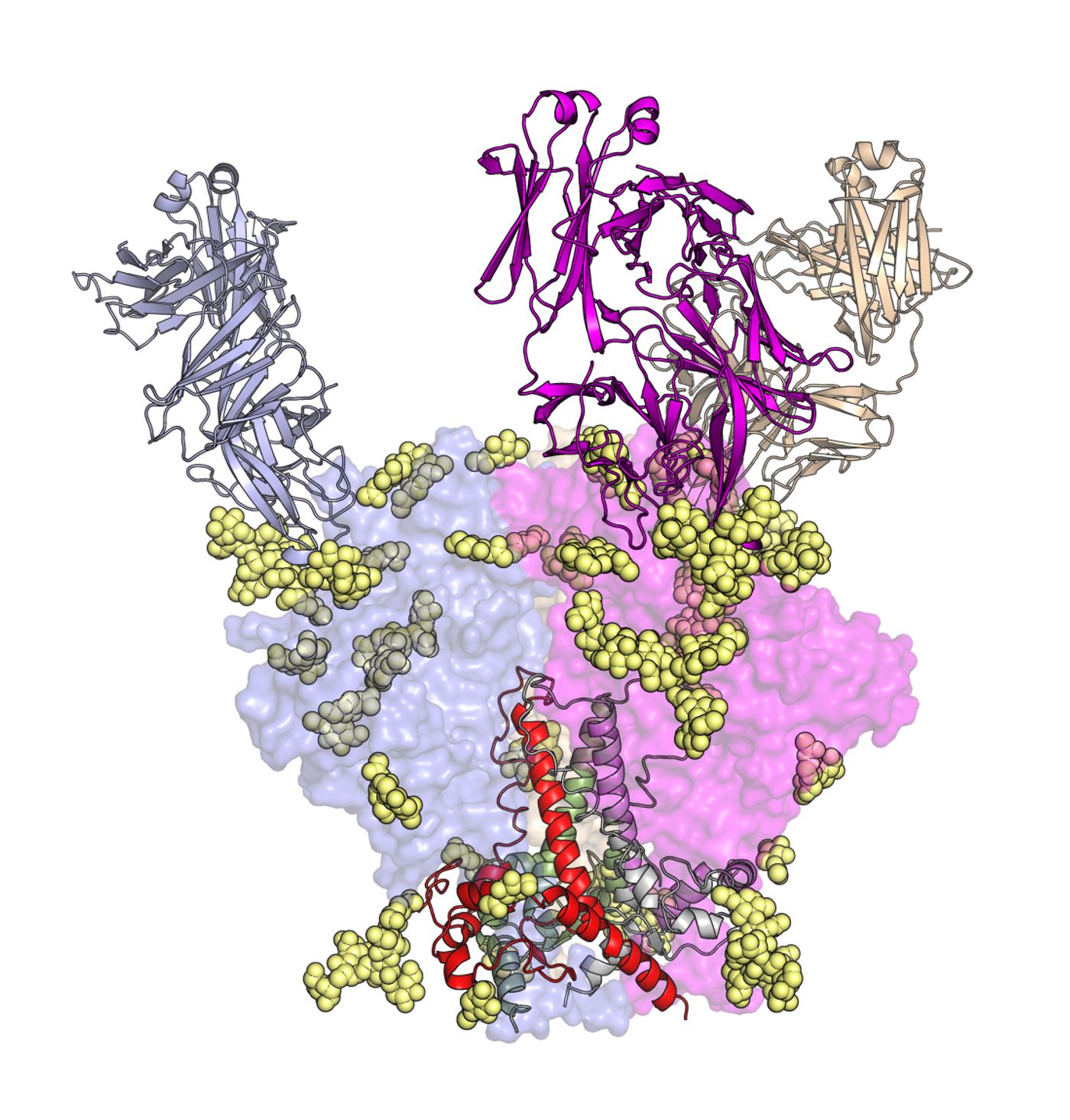
Using high-resolution imaging techniques, the researchers mapped out the structures of these antibodies, bound with the HIV Envelope glycoprotein, at select points in their boot-camp training. They found that different branches of antibodies developed different strategies to get around HIV’s defenses.
The Anti-HIV Arms Race
The antibodies’ main obstacle is HIV’s glycan shield, a layer of sugars that sits on the HIV Envelope glycoprotein, protecting it from the body’s antibody attacks.
“The virus is always trying to escape the immune system, and the immune system—via antibodies—tries to go after it,” said Fernando Garces, research associate at TSRI and first author of the new study. “What makes PGT121 family of antibodies so special,” he added, “is that it has found ways to counter-attack the virus while many other families of antibodies fail to do so.”
The images in the new study showed that the PGT121 family of antibodies starts its attack by grabbing onto one of the glycans, (called the N332 glycan) as well as a short segment of the HIV Envelope glycoprotein (called the GDIR motif), turning them into binding partners.
From there, the researchers observed that antibodies within this family developed slightly different strategies. While antibodies from the PGT122 branch mutate to engage other surrounding glycans (the N137 glycan), the PGT124 branch appears to do the same, but then circumvents other elements of the glycan shield (again, the N137 glycan) by rotating and tilting to target the HIV Envelope glycoprotein beneath.
Working Toward a Rapid Response
Interestingly, if researchers remove the N137 glycan from HIV, the antibodies immediately behave as well-trained antibodies and don’t require a two-year boot camp.
“That suggests possible strategies for how one might elicit those antibodies through vaccination,” said Wilson.
If scientists can design a molecule that lacks this N137 glycan, they may be able to prompt a faster antibody response. Once the antibody response has been kick-started, they could add this glycan back and try to mature the antibody against the more native form of the virus.
The new study also showed a much-improved 3D image of the HIV Envelope protein. This detailed “map” of the final pieces of HIV’s trimer structure will help researchers design HIV vaccines.
“The take-away message is that you’ve really got to track the evolution of the antibody and the antigen [the target on a virus] in order to understand how the immune response targets HIV,” said Wilson.
Wilson added that the study required a “team approach.” The findings resulted from collaborations with the laboratory of TSRI Professor Dennis R. Burton, who is also scientific director of the International AIDS Vaccine Initiative (IAVI) Neutralizing Antibody Center and of the National Institutes of Health (NIH) Center for HIV/AIDS Vaccine Immunology and Immunogen Discovery (CHAVI-ID) at TSRI, which produced the antibodies for the study; the laboratory of TSRI Associate Professor Andrew Ward, which performed the electron microscopy imaging that helped reveal how the antibodies bind; and the labs of Rogier Sanders of the University of Amsterdam and Weill Medical College of Cornell University and John P. Moore of Weill Medical College of Cornell University.
In addition to Wilson, Garces, Burton, Ward, Sanders and Moore, authors of the study, “Affinity maturation of a potent family of HIV antibodies is primarily focused on accommodating or avoiding glycans,” were Jeong Hyun Lee, Natalia de Val, Leopold Kong, Robyn L. Stanfield of TSRI, IAVI and CHAVI-ID; Cristina Puchades and Yuanzi Hua of TSRI; and Alba Torrents de la Pena of the University of Amsterdam.
The study was supported by the U.S. Department of Energy (contracts DEAC02- 05CH11231 and DE-AC02-06CH11357), the HIV Vaccine Research and Design (HIVRAD) program (grants P01 AI082362, P01 AI110657 and R37 AI036082), the Center for HIV/AIDS Vaccine Immunology and Immunogen Discovery (CHAVI-ID UM1 AI00663), IAVI Neutralizing Antibody Center, NIH (grants RO1 GM046192 and 2R56 AI084817 ), the Joint Center of Structural Genomics funded by the NIH National Institute of General Medical Sciences (NIGMS) Protein Structure Initiative (grant U54 GM094586), the Netherlands Organization for Scientific Research, the European Research Council (ERC-StG-2011–280829-SHEV), and the California HIV/AIDS Research Program Dissertation Award (D12-SRI-353).
_________
FOR MORE INFORMATION
Ian Wilson Biosketch
Wilson Lab Website
Immunity
This article was originally published on
scripps.edu