Database for Functional Plant Microbiome Studies
Humanity will need to find solutions to feed a growing population; there are currently 7.6 billion people on the planet, and the number is expected to hit ten billion by 2050. The relationships between microbes and plants may provide new ways to improve agricultural yield in the face of environmental challenges like drought, make use of non-arable land, or reduce the impact on the environment caused by farming. To aid researchers who are investigating the community of microbes that live among plants and how they interact, scientists have identified genes that are likely candidates for assisting bacteria in the adaptation to the plant environment.
Previous work on the plant microbiome has focused on identifying the members of the bacterial community, and individual relationships between a bacterial strain and a plant. This work investigates genes that play a role in bacterial root colonization. It has been reported in Nature Genetics by scientists at the U.S. Department of Energy (DOE) Joint Genome Institute (JGI), a DOE Office of Science User Facility, and the Howard Hughes Medical Institute at the University of North Carolina at Chapel Hill (UNC).
"If we want to engineer the right microbiome to support plant growth, we need to understand the real function of the microbiome and not just sequence marker genes," explained the co-first author of the work Asaf Levy, a research scientist at the JGI. "Here we used a massive genomic and computational effort to address the fundamental and important question: 'How does the plant microbiome interact with the plant?'"
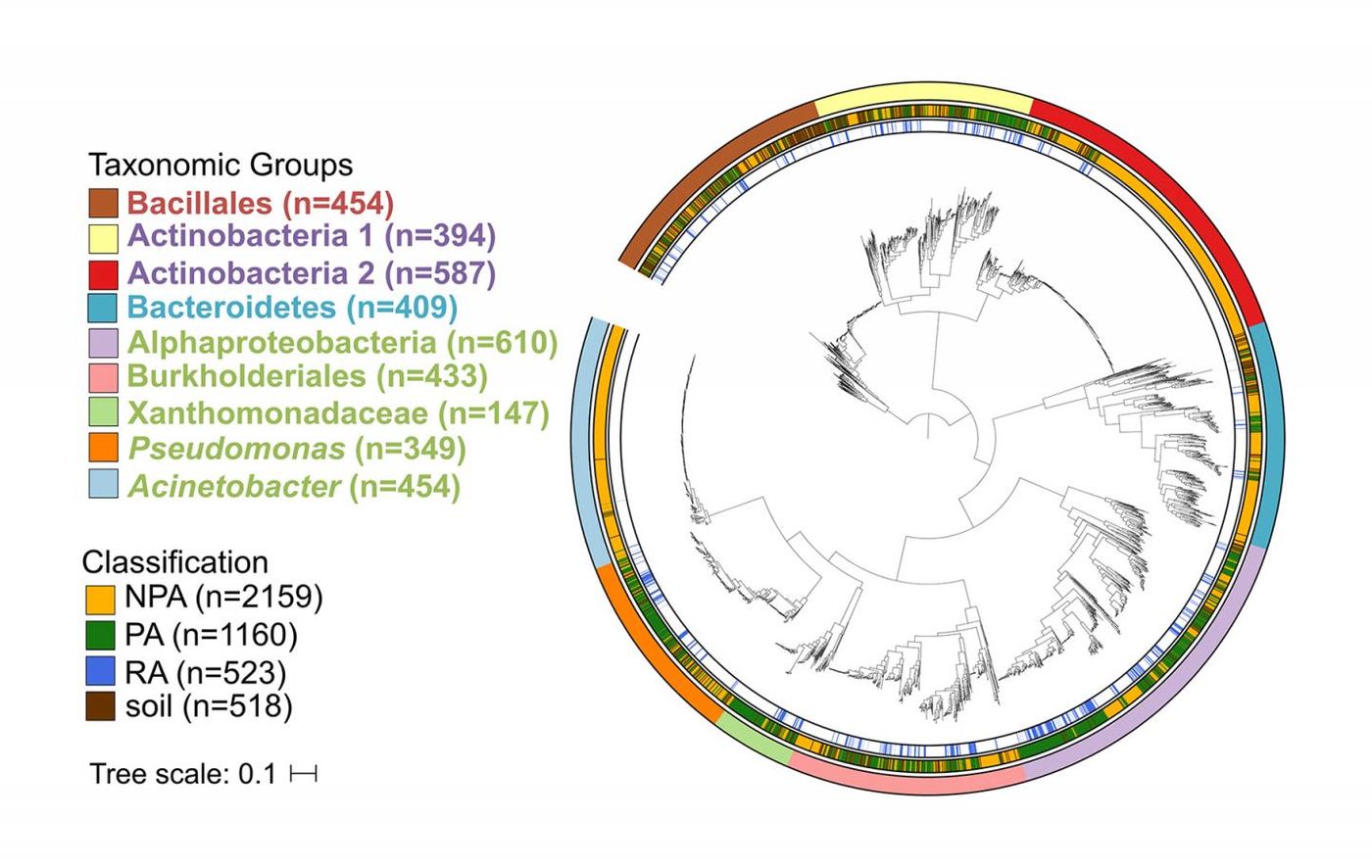
Most microbes interact with plants where the roots meet the soil. The researchers analyzed the genomes of 377 novel bacterial isolates taken from maize (51), poplar trees (135), Brassicaceae (191), in addition to 107 single bacterial cells taken from A. thaliana. The data were compared with available genomic information representing the main groups of bacteria that have been associated with plants as well as bacteria from totally different sources, like the human gut. This allowed the investigators to identify genes that are enriched in the genomes of microbes that are associated with plants, or not.
"It's very important for us to understand what genes and functions microbes use to colonize plants because only then might we have a chance to rationally devise useful 'plant probiotics' to help us raise more food and energy crops with fewer chemical inputs such as fertilizers and pesticides or fungicides," explained the senior author of the study Jeff Dangl, a Howard Hughes Medical Institute investigator and the John N. Couch Professor of Biology at the University of North Carolina at Chapel Hill.
The scientists found that genes associated with the metabolism and transport of sugar are more common in plant- and soil-associated genomes, which tends to make them larger. Because of photosynthesis, plants secrete carbon from their roots as sugars, attracting microorganisms.
The researchers also found numerous genes that appear to mimic plant functions; they encode "Plant-Resembling PA and RA Domains" or PREPARADOs. "It is well known that plant pathogens use proteins that mimic plant domains required for immune function," said Dangl. "Imagine that the pathogen injects directly into the plant cell a protein that mimics part of a particular immune system machine. It's like putting a partly defective cog into a wheel--the wheels can't turn anymore. We reckon that the plant-associated protein domains that we identified might work in the same way."
A complete catalog of genes associated with plants and new genome sequences are now open to the research community through Genomic Features of Bacterial Adaptation to Plants. Laboratories interested in collaborating with the Berkeley Lab should check out IPO's Industry Partnerships and email ipo@lbl.gov.
"The database is a precious resource for the research community studying plant-microbe interactions as it is an unbiased way to identify potentially interesting genes involved in interaction with a plant--including many totally novel genes. We are currently experimentally studying the function of many of these genes to gain a better functional understanding of the plant microbiome." Levy concluded.
Sources: AAAS/Eurekalert! Via DOE/Joint Genome Institute, Nature Genetics