Looking to Bacteria for the Next Antibiotic
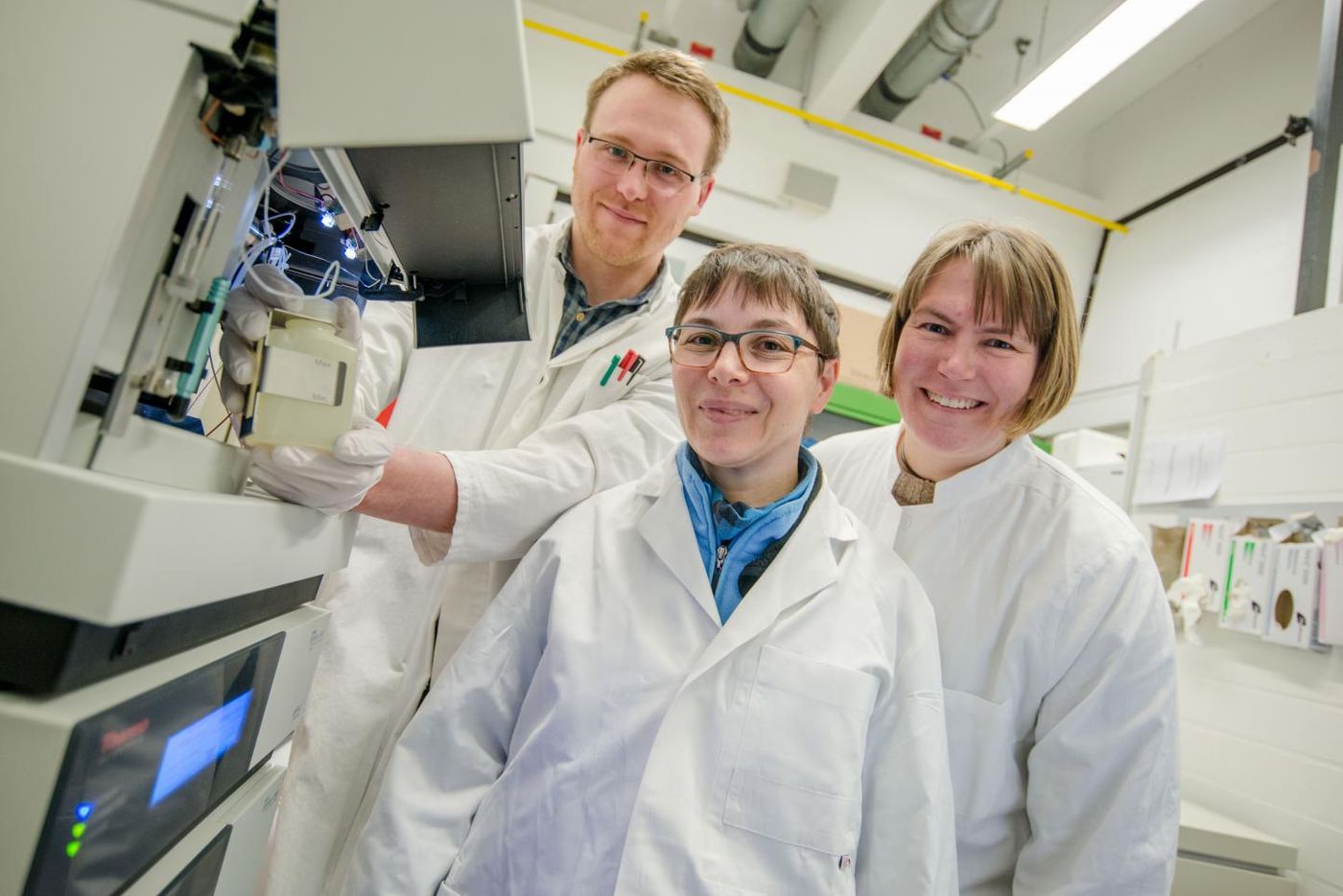
Bacteria can be a surprising source of antibiotics. One example is Streptomyces chartreusis, which can secrete a variety of compounds into its environment, more than had been assumed. Many of these bacterial molecules have potential as pharmacological agents. A team of researchers led by Professor Julia Bandow and Christoph Senges of the Applied Microbiology research group at Ruhr-Universität BochumX and an international group of collaborators reported their findings in the Proceedings of the National Academy of Sciences.
Various Streptomyces bacteria make around 70 percent of all the natural antibiotics used in the clinic to treat people. The scientists found that the Streptomyces chartreusis genome carries 128 gene clusters which may be important to the production of molecules. "This is a fair number, but the number of secreted molecules that we detected exceeded our expectations," revealed Bandow.
The researchers provided three different kinds of environmental compositions for the bacteria to grow in and harvested the compounds they secreted. They found 1,044 substances. Different conditions caused the production of different molecules as well; some stuff was only made, for example, if no iron was present.
Eight new molecules that are critical for the uptake of iron, desferrioxamines, were found; that class of compounds helps treat patients who suffer overdoses of aluminum or iron. "This shows the potential of detecting new variants of well-known medicines," explained Bandow.
The structures of the remaining bacterial chemicals are primarily unknown. The scientists expect that their bacteria have created totally new kinds of molecules. "Based on our results, we anticipate that many aspects of bacterial chemistry including chemical structures, pharmacological potential, and ecological significance are yet to be uncovered," noted Bandow.
The team also showed that a biosynthetic gene cluster can be induced to make a range of chemicals. “Most likely, this phenomenon reflects an adaptation of the organism to different conditions," said Christoph Senges.
For this work, a specialized technique called tandem mass spectrometry was utilized to compare the mixtures of chemicals made naturally and secreted by the bacteria into their cultures; it also found substances that were only present in tiny amounts. The method also enabled the prediction of chemical structures for the novel compounds.
Learn more about how bacteria actually make antibiotics and the gene sequences associated with some of those drugs, from this video. After an overview of how DNA is made into proteins, the rest of the talk moves on to antibiotic production around the four-minute mark.
Sources: AAAS/Eurekalert! Via Ruhr-Universität Bochum, PNAS