Junk DNA refers to regions of the genome that did not at first appear to have any function or relevance. Research over time has revealed that in fact, these regions have importance, even if it’s not well understood. Long noncoding RNA (lncRNA) refers to the RNA copied from an area of the genome thought of as junk DNA. It’s been shown that lncRNA has an influence on cell fate as the embryo develops, although how that is accomplished is unknown. Want to know more about noncoding RNA? Watch the video below.
Biologists at MIT, inspired by classical work on RNA that shows how structure is intimately related to function, took a closer look at the structure of lncRNA. They have linked the structure of one type of lncRNA to its function in
new work published in Molecular Cell.
"Emerging data points to fundamental roles for many of these molecules in development and disease, so we believe that determining the structure of lncRNAs is critical for understanding how they function," explained Laurie Boyer, the senior author of the study and the Irwin and Helen Sizer Career Development Associate Professor of Biology and Biological Engineering at MIT.
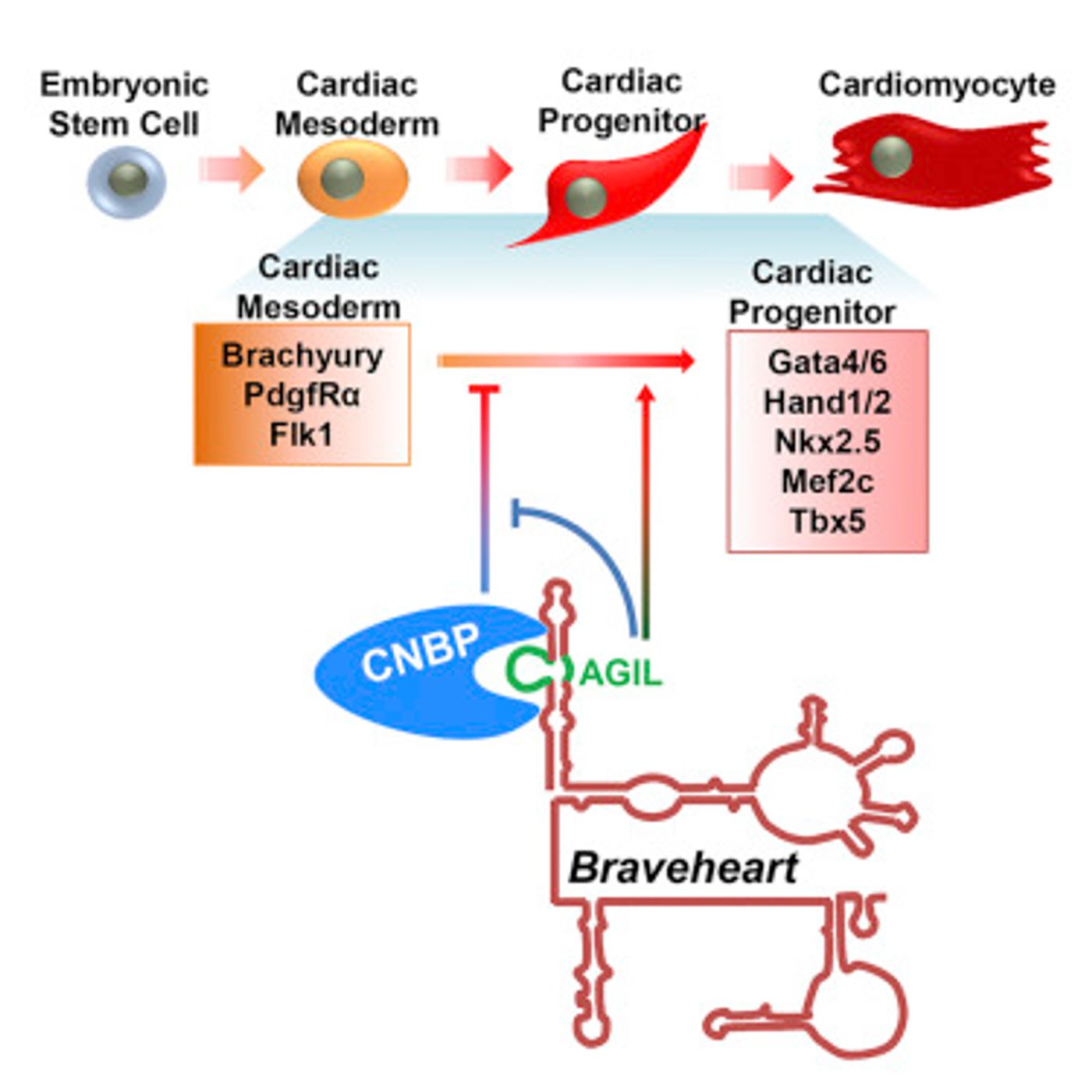
Boyer’s lab has previously demonstrated that a mouse lncRNA which is called Braveheart and is found at high levels in heart tissue, is required for the normal development of muscle cells of the heart.
In the new work, the research team wanted to determine which parts of the RNA molecule were necessary for its function. "We knew Braveheart was critical for heart muscle cell development, but we didn't know the detailed molecular mechanism of how this lncRNA functioned, so we hypothesized that determining its structure could reveal new clues," explained lead author Zhihong Xue, an MIT postdoc.
Using a method called chemical probing, the scientists investigated the structure. The RNA molecule is treated with a chemical that modifies exposed nucleotides. As such, identifying features like loops, single or double stranded regions, and helixes in the molecule can be revealed.
It was found that Braveheart has several motifs or structural regions. Further analysis showed that the removal of 11 particular nucleotides of the molecule was enough to stop normal development of heart cells. The researchers then looked for what proteins could interact with that 11 nucleotide loop, and found cellular nucleic acid binding protein (CNBP), a transcription factor that binds strongly to this region. Additionally, studies have shown that CNBP mutations can cause heart defects in both mice and humans.
Work showed that CNBP can act as a brake on heart development, and Braveheart releases the brake, resulting in heart muscle development.
While a human ortholog to Braveheart has not yet ben found, the investigators plan to search for similar structures in human lncRNAs.
"We're taking this motif and we're using it to build a fingerprint so we can potentially find motifs that resemble that lncRNA across species," Boyer continued. "We also hope to extend this work to identify the modes of action of a catalog of motifs so that we can better predict lncRNAs with important functions."
The researchers also hope to use what they have learned about lncRNA for the development of new therapeutics. "We fully expect that unraveling lncRNA structure-to-function relationships will open up exciting new therapeutic modalities in the near future," Boyer concluded.
If you’d like to know more about Boyer’s research, watch her 30 minute lecture given at the 2014 Koch Institute Summer Symposium at MIT, shown in the video above.
Sources:
AAAS/Eurekalert! via
MIT,
Molecular Cell